Population genetics of rapid adaptation and the predictability of evolution
Richard Neher
Biozentrum, University of Basel
slides at neherlab.org/201706_facmtg.html
- Evolution of HIV
- Theory of evolution
- Tracking and prediction of RNA virus evolution
Evolution of HIV
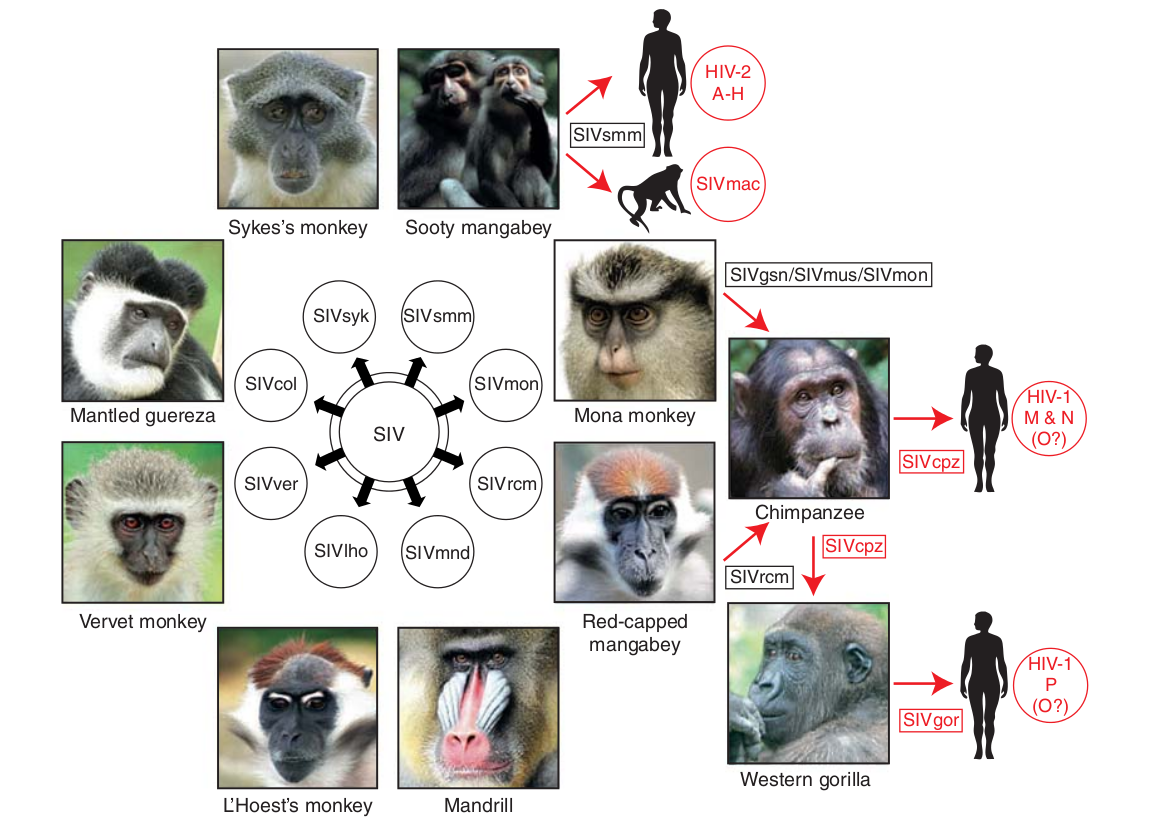
- Chimp → human transmission ~1900 gave rise to HIV-1 group M
- Diversified into subtypes that are ~20% different
- evolves at a rate of about 0.1% per year
HIV-1 sequencing before and after therapy
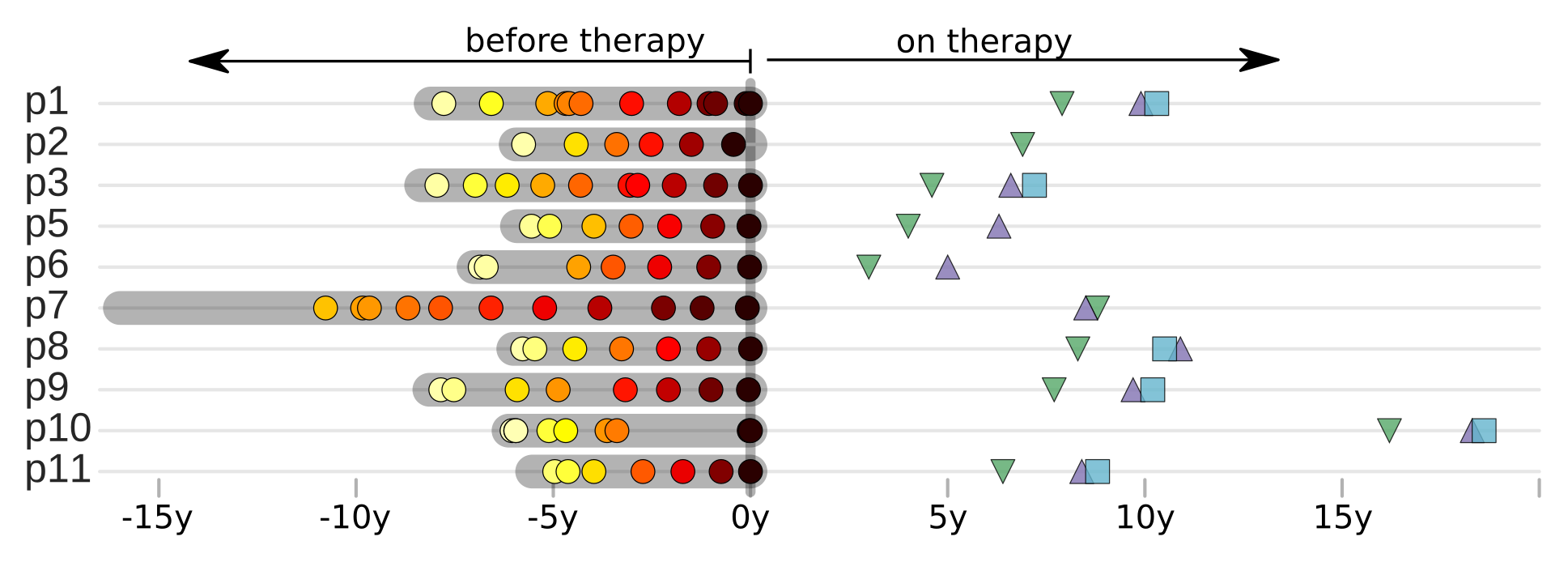 Zanini et al, eLife, 2015;
Brodin et al, eLife, 2016.
Collaboration with the group of Jan Albert
Zanini et al, eLife, 2015;
Brodin et al, eLife, 2016.
Collaboration with the group of Jan Albert
Population sequencing to track all mutations above 1%
- diverge at 0.1-1% per year
- almost full genomes coverage in 10 patients
- full data set at hiv.tuebingen.mpg.de
Rapid evolution before therapy, no evolution on therapy
Population genetics models
evolutionary processes ↔ trees ↔ genetic diversity
Kingman coalescent
strong selection
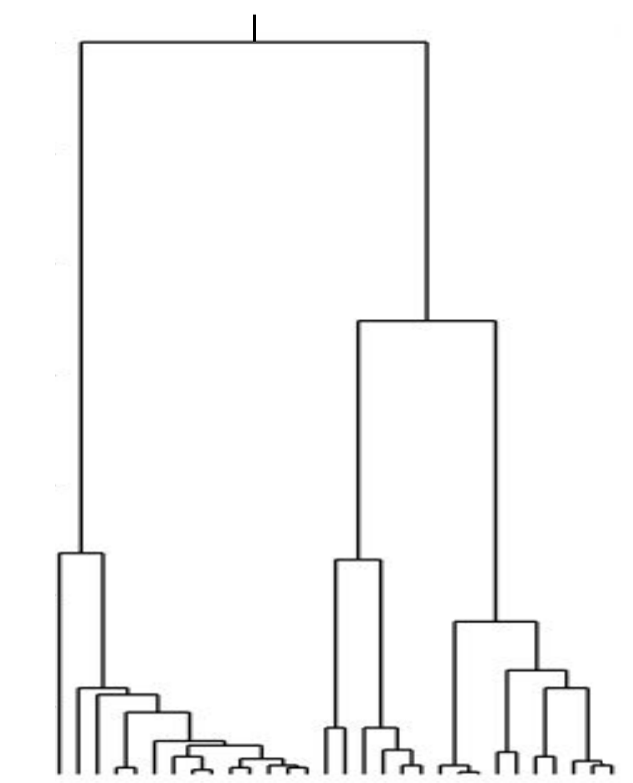
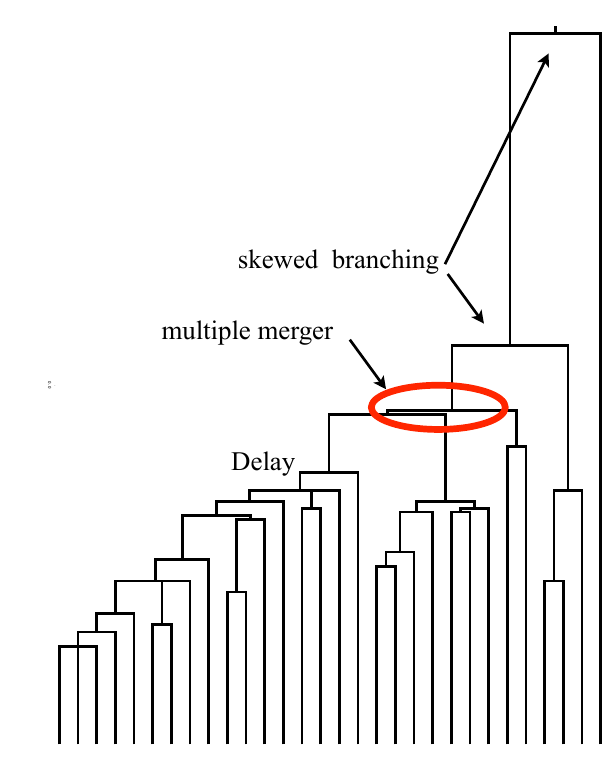
Bolthausen-Sznitman Coalescent
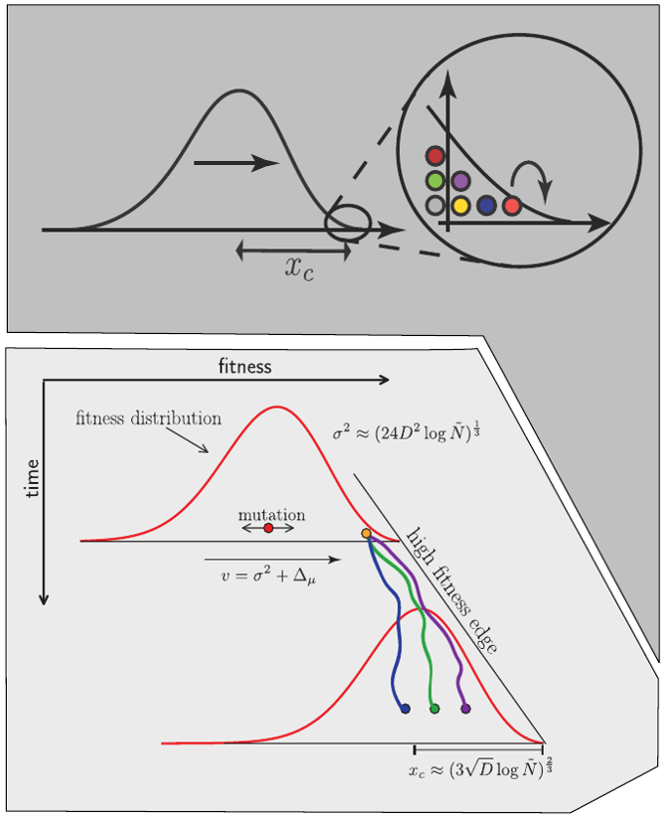
Bursts in a tree ↔ high fitness genotypes
Can we read fitness of a tree?



- Influenza virus evolves to avoid human immunity
- Vaccines need frequent updates

Fitness inference from trees
$$P(\mathbf{x}|T) = \frac{1}{Z(T)} p_0(x_0) \prod_{i=0}^{n_{int}} g(x_{i_1}, t_{i_1}| x_i, t_i)g(x_{i_2}, t_{i_2}| x_i, t_i)$$
RN, Russell, Shraiman, eLife, 2014
Prediction of the dominating H3N2 influenza strain
RN, Russell, Shraiman, eLife, 2014nextflu.org
nextstrain.org
HIV acknowledgments
- Fabio Zanini
- Jan Albert
- Johanna Brodin
- Christa Lanz
- Göran Bratt
- Lina Thebo
- Vadim Puller



Influenza and Theory acknowledgments
- Boris Shraiman
- Colin Russell
- Trevor Bedford


