Population genetics of rapid adaptation and the predictability of evolution
Richard Neher
MPI for Developmental Biology
slides at neherlab.org/201701_vienna.html
Evolution of HIV
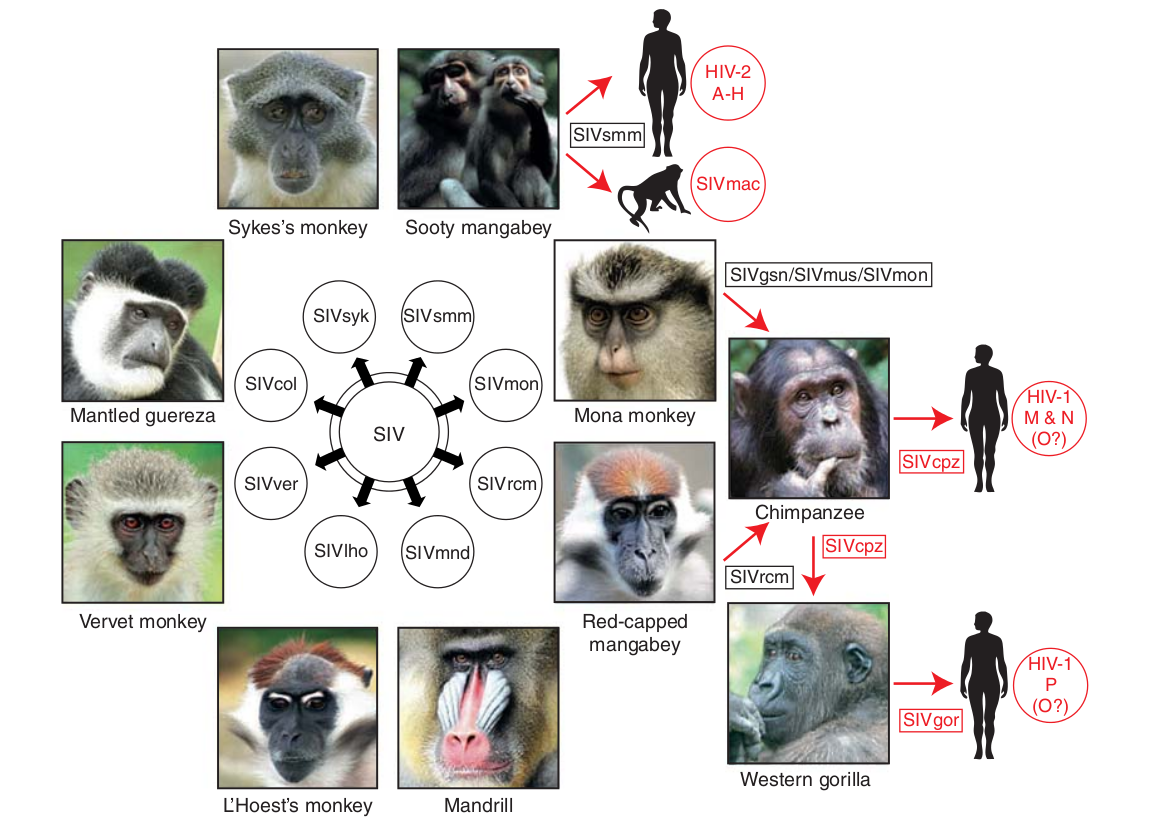
- Chimp → human transmission ~1900 gave rise to HIV-1 group M
- Diversified into subtypes that are ~20% different
- evolves at a rate of about 0.1% per year
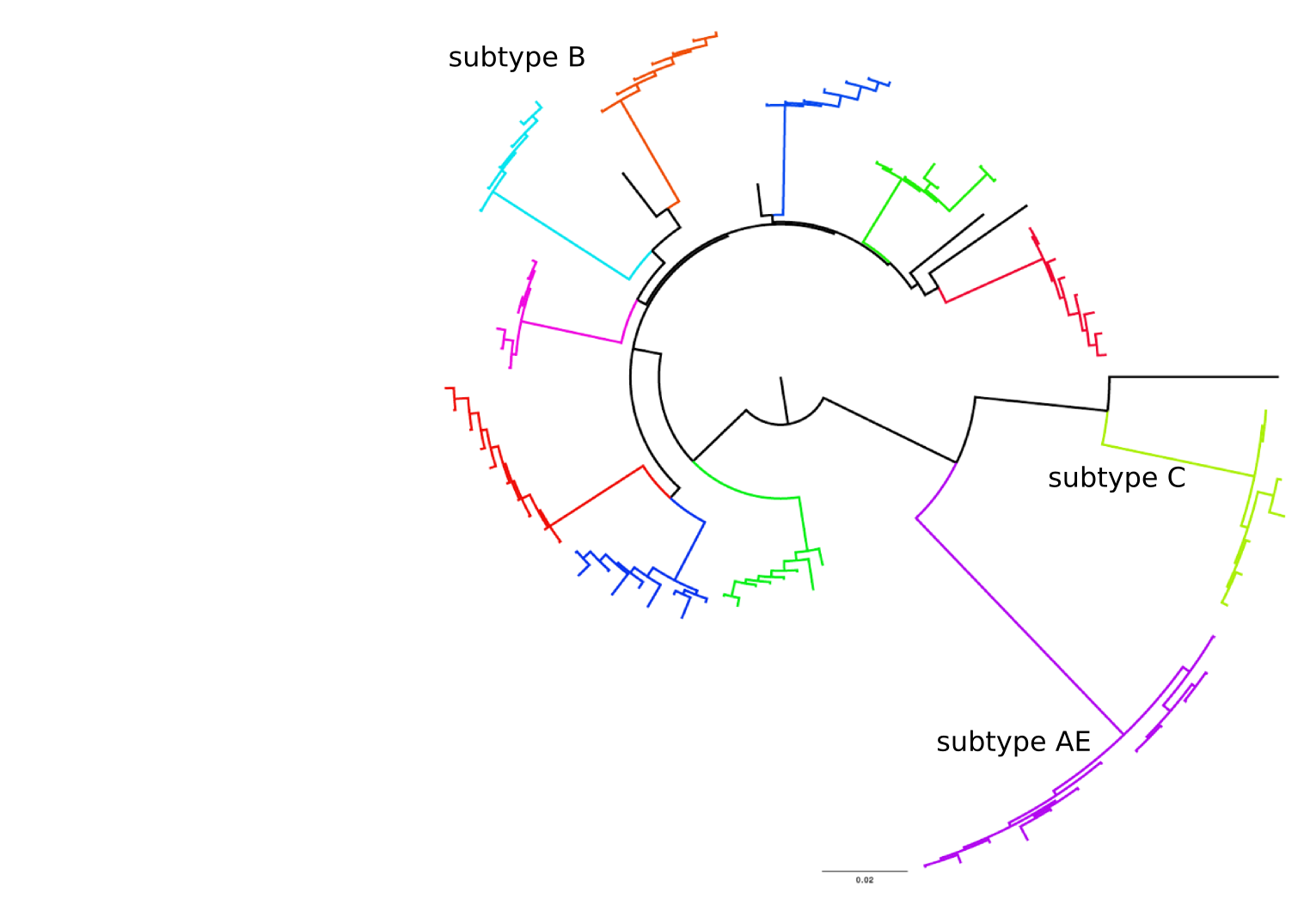
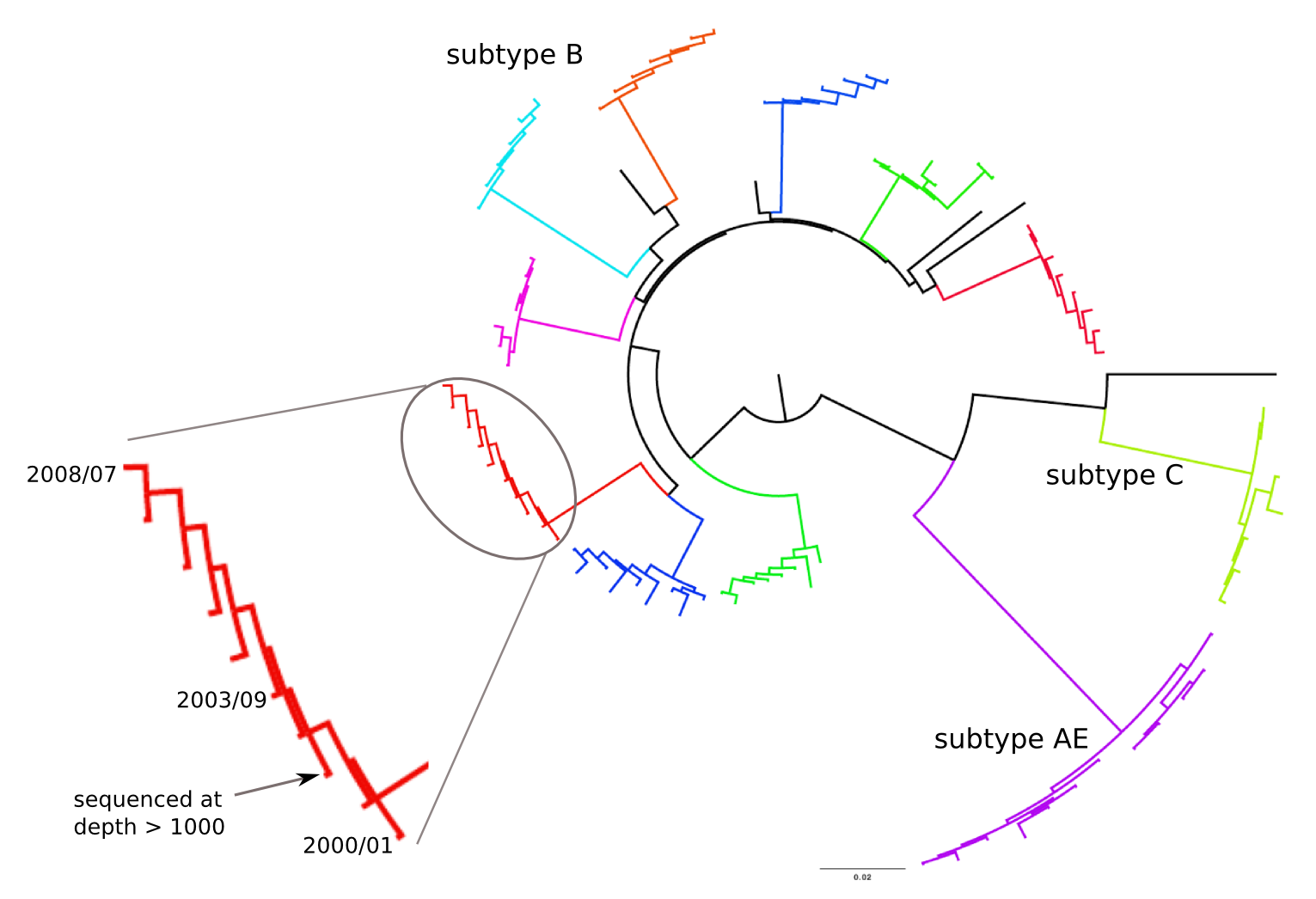
Population sequencing to track all mutations above 1%
- diverge at 0.1-1% per year
- almost full genomes coverage in 10 patients
- full data set at hiv.tuebingen.mpg.de
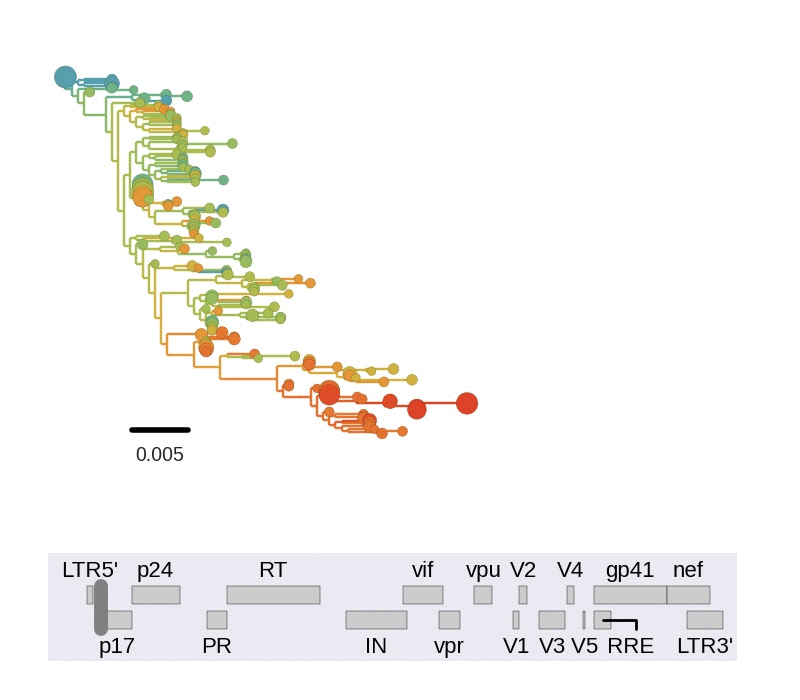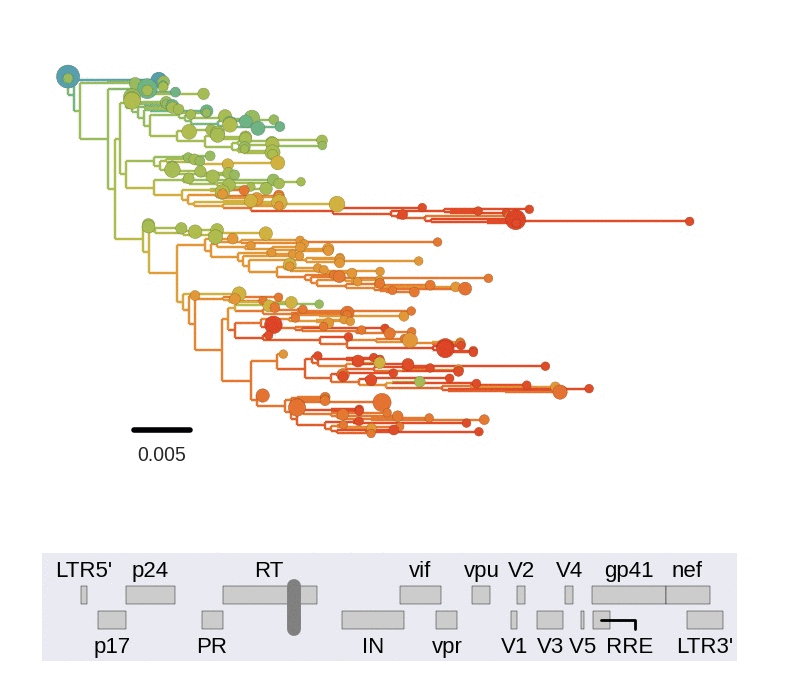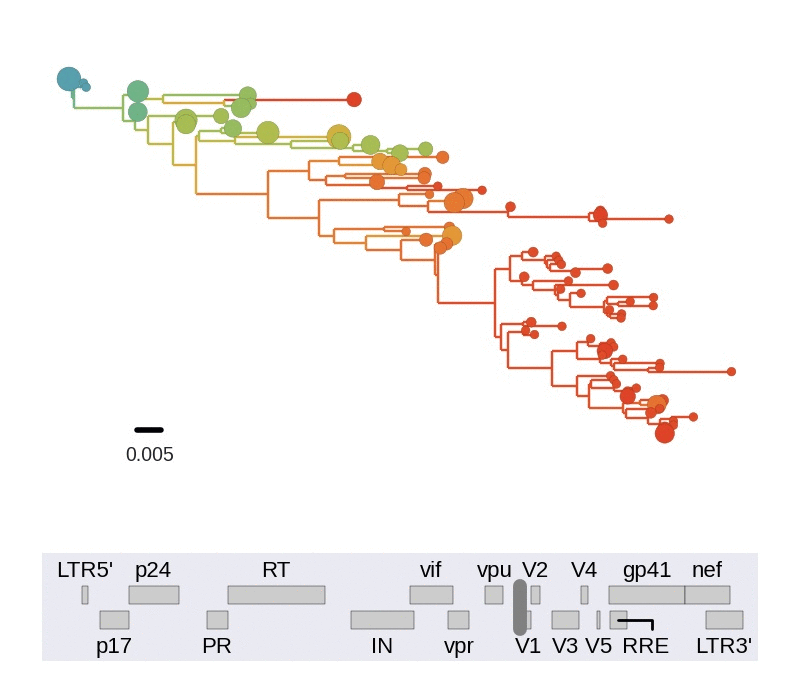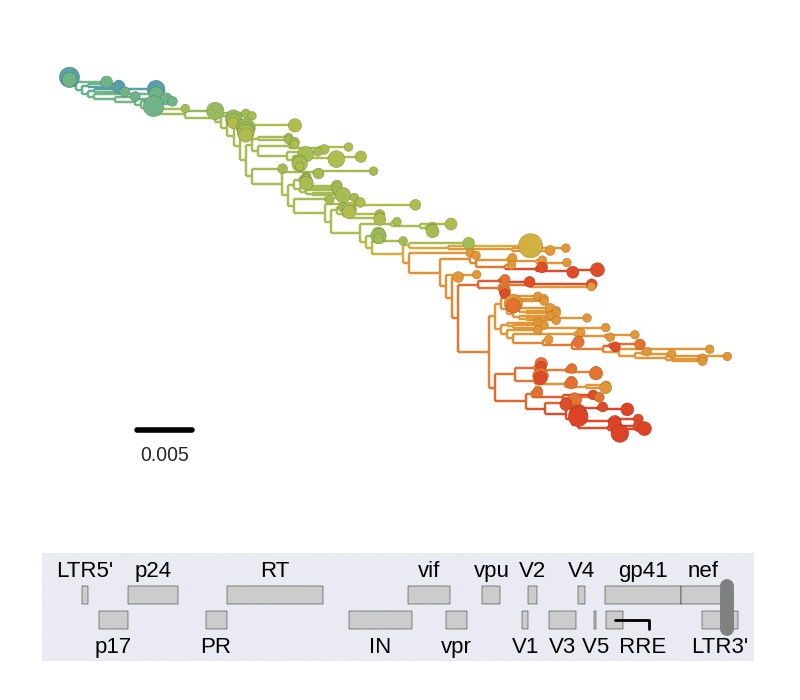
Diversity and hitchhiking
- envelope changes fastest, enzymes lowest
- identical rate of synonymous evolution
- diversity saturates where evolution is fast
- synonymous mutations stay at low frequency
Frequent version of previously beneficial mutations
- HIV escapes immune systems
- most mutations are costly
- humans selects for different mutations
- are there costs compensated or reverted
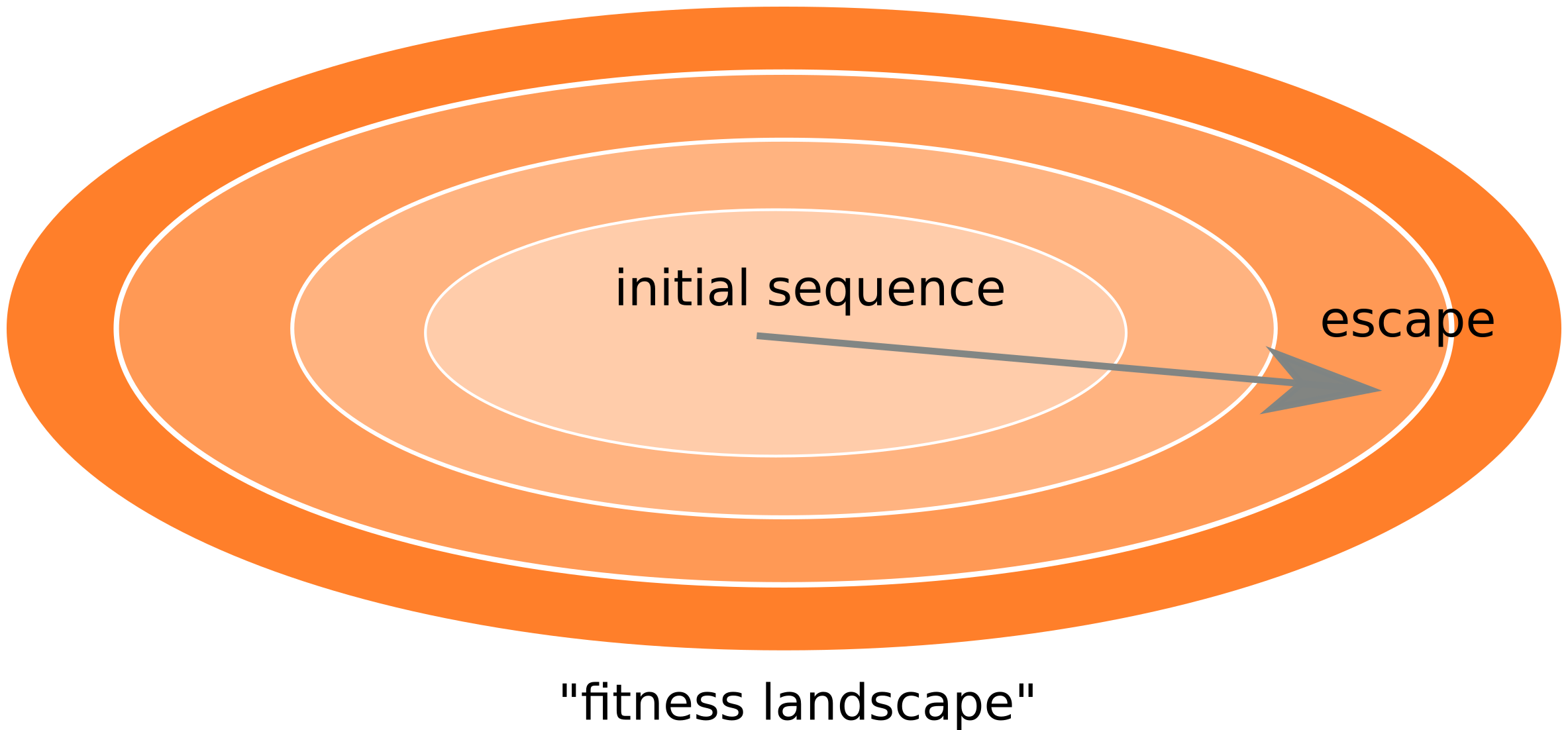
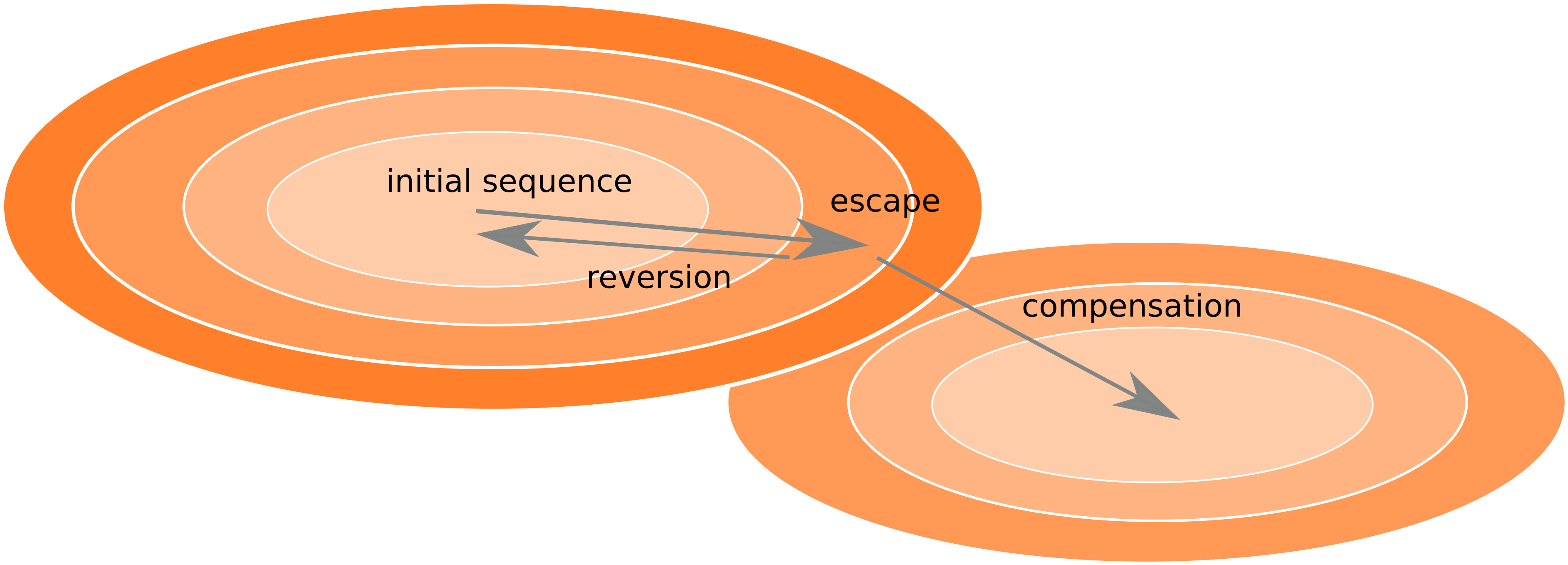
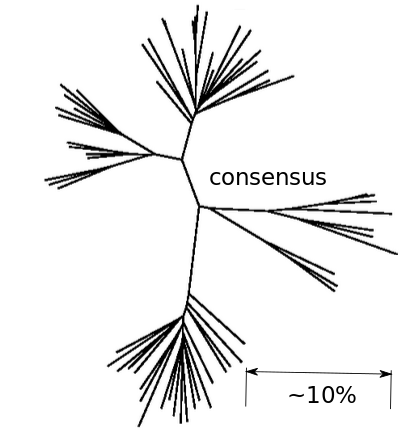
- Almost one third of all mutations are reversions vs expected: 5%
- Strong evolutionary attractor
Fitness landscape of HIV-1
Zanini et al, biorxiv, 2017Selection on RNA structures and regulatory sites
Zanini et al, biorxiv, 2017The distribution of fitness effects
Zanini et al, biorxiv, 2017Population genetics models
evolutionary processes ↔ trees ↔ genetic diversity
Neutral models and beyond
Neutral models
- all individuals are identical → same offspring distribution
- Kingman coalesence and diffusion theory are dual descriptions
- everything is easy to calculate
- perturbations like background selection can be included
What if selection is everywhere?
Clonal interference and traveling waves
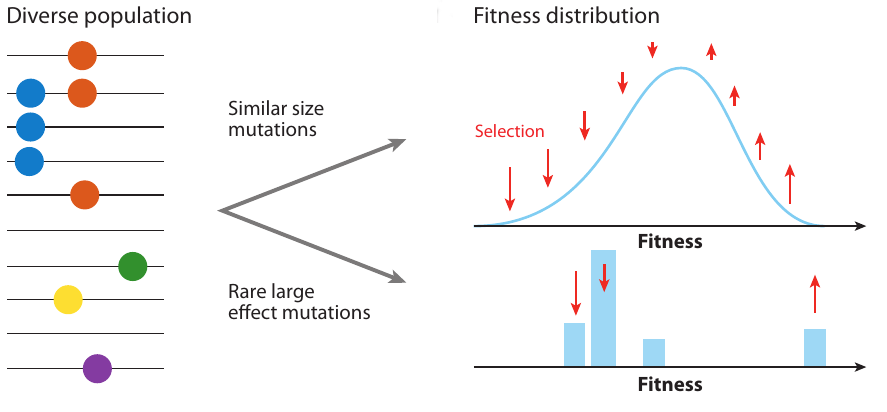
- extensive work on speed of adaptation, but this speed is not observable
- genetic diversity is what we observe
- depends on the properties of trees
What if selection is everywhere?
Kingman coalescent
strong selection
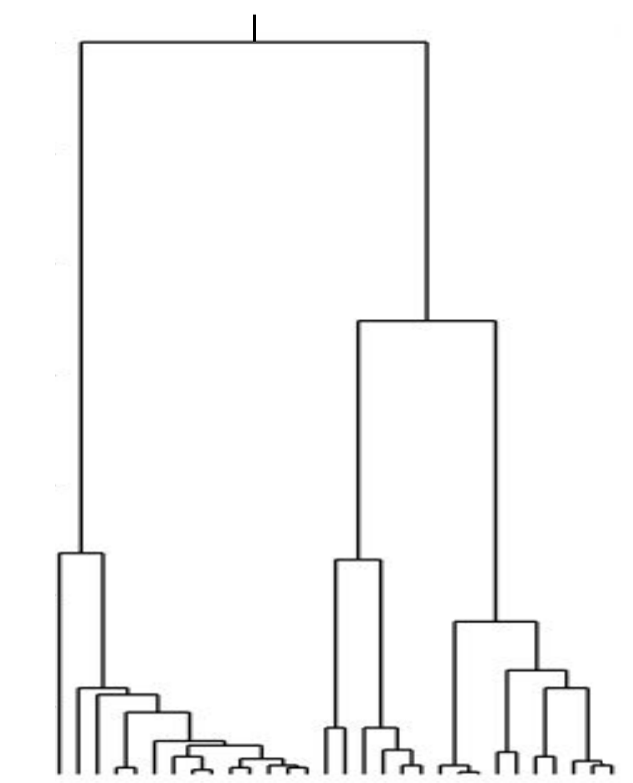
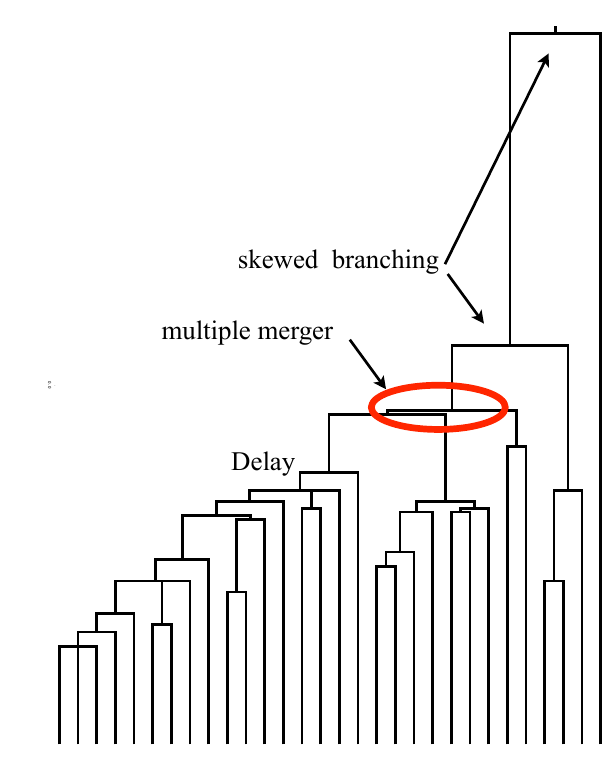
Bolthausen-Sznitman Coalescent
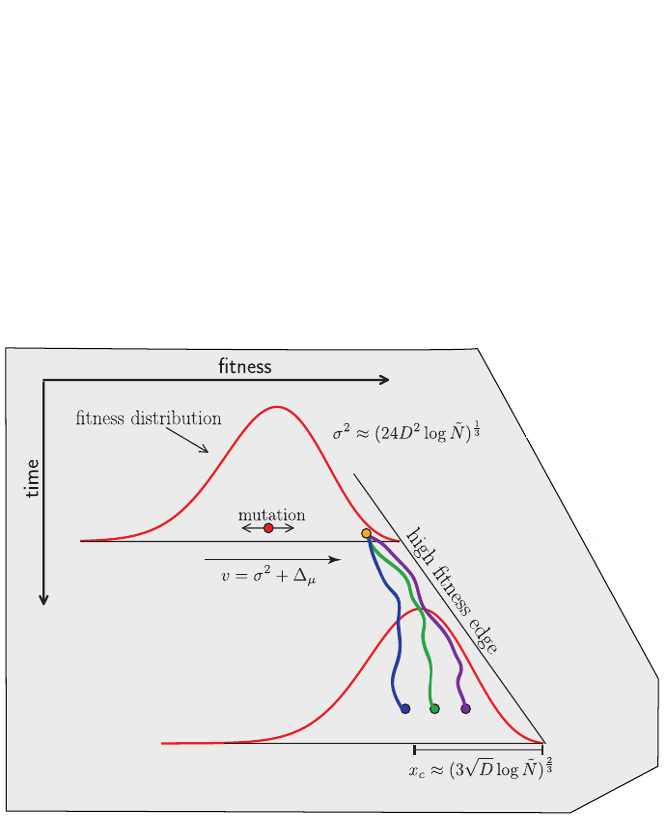
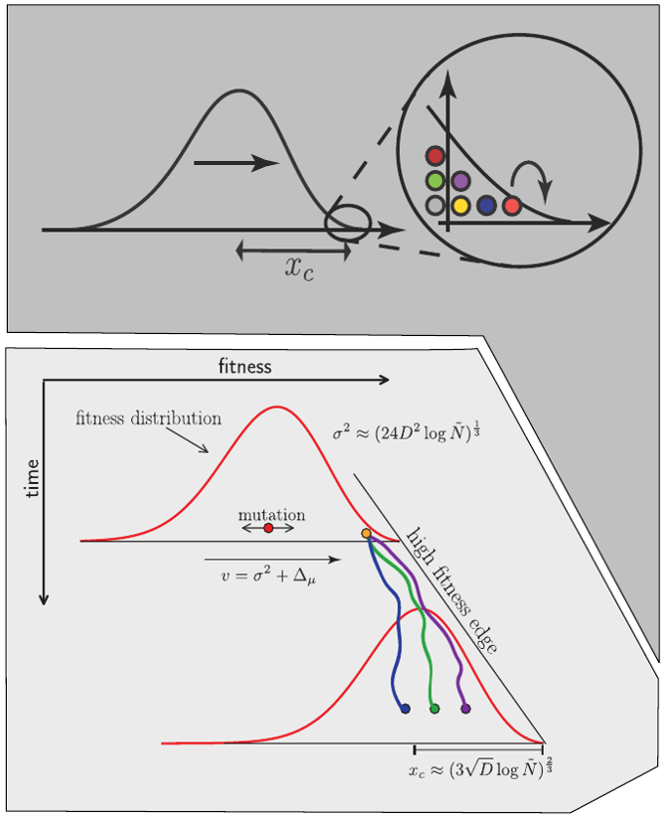
Universality of the Bolthausen-Sznitman Coalescent (BSC)
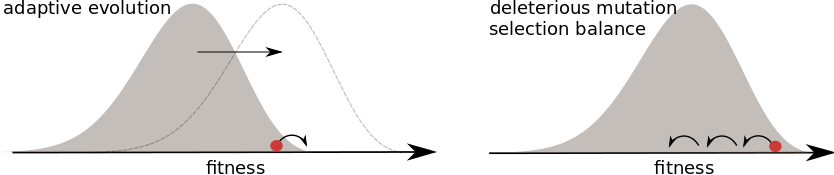
- many small effect mutations → coalescence is BSC like
- fitness diversity $\sigma$, not population size determines $T_{MRCA}$
- the time scale of coalescence is always $T_c \sim \sigma^{-1}\sqrt{\log N}$
- frequency dynamics is not diffusive, but has Levy-flight properties
- Can be extended to sexual populations
U-shaped polarized site frequency spectra
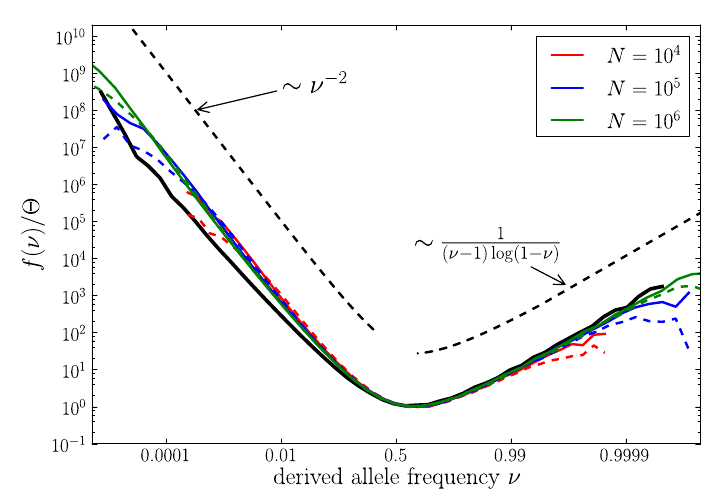

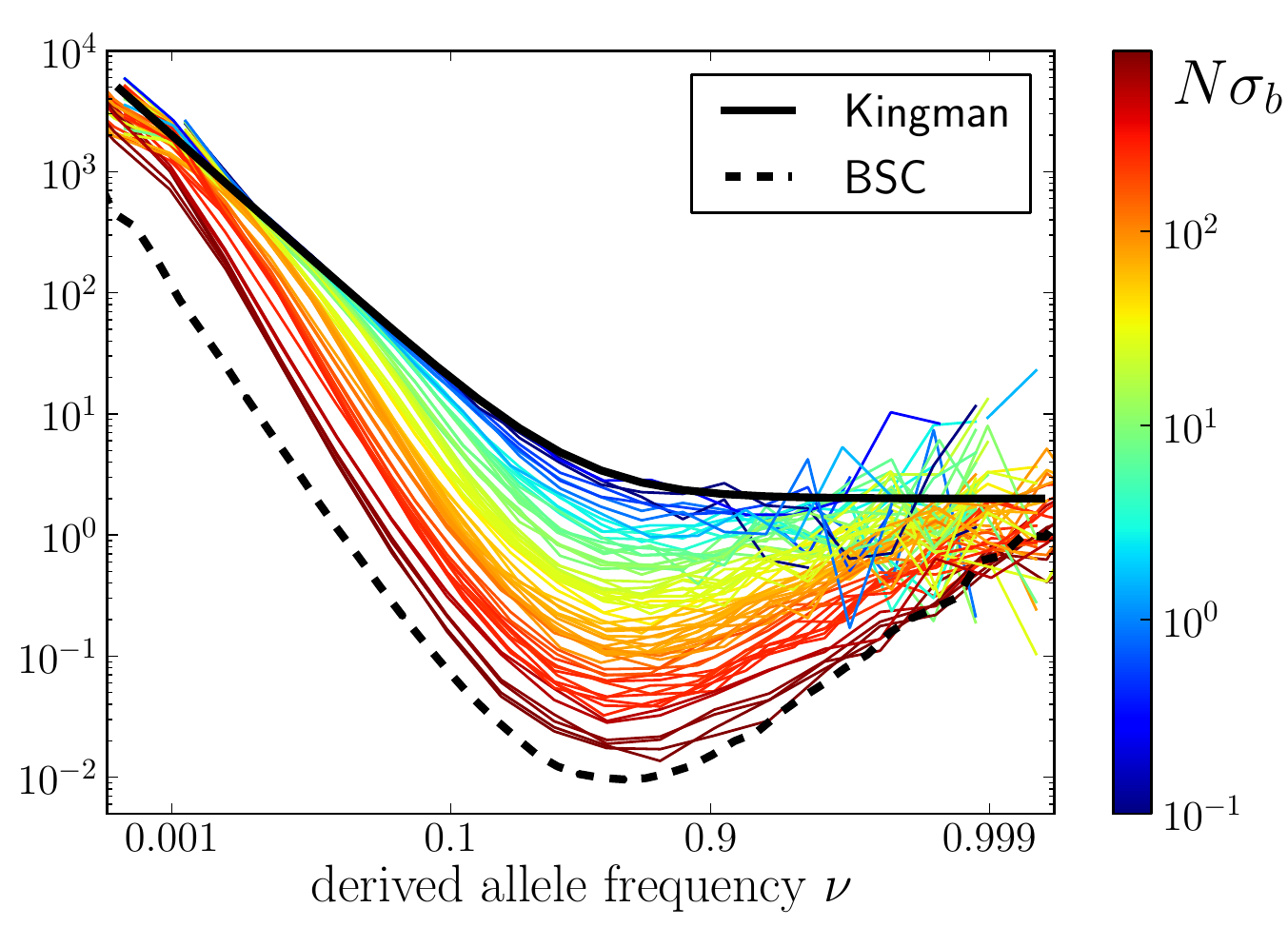 RN, Kessinger, Shraiman, PNAS, 2013
RN, Kessinger, Shraiman, PNAS, 2013
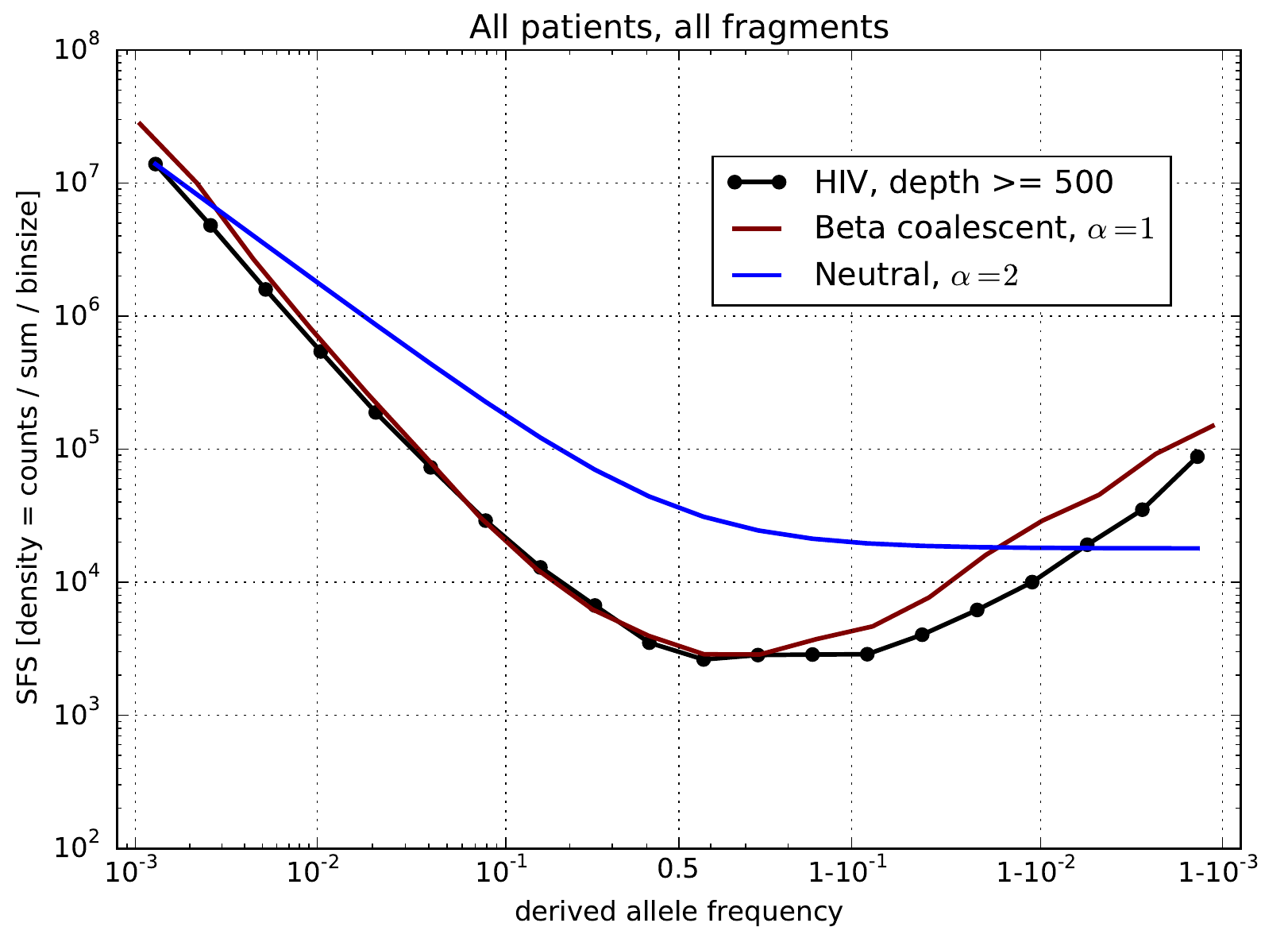 Zanini et al, eLife, 2015
Zanini et al, eLife, 2015
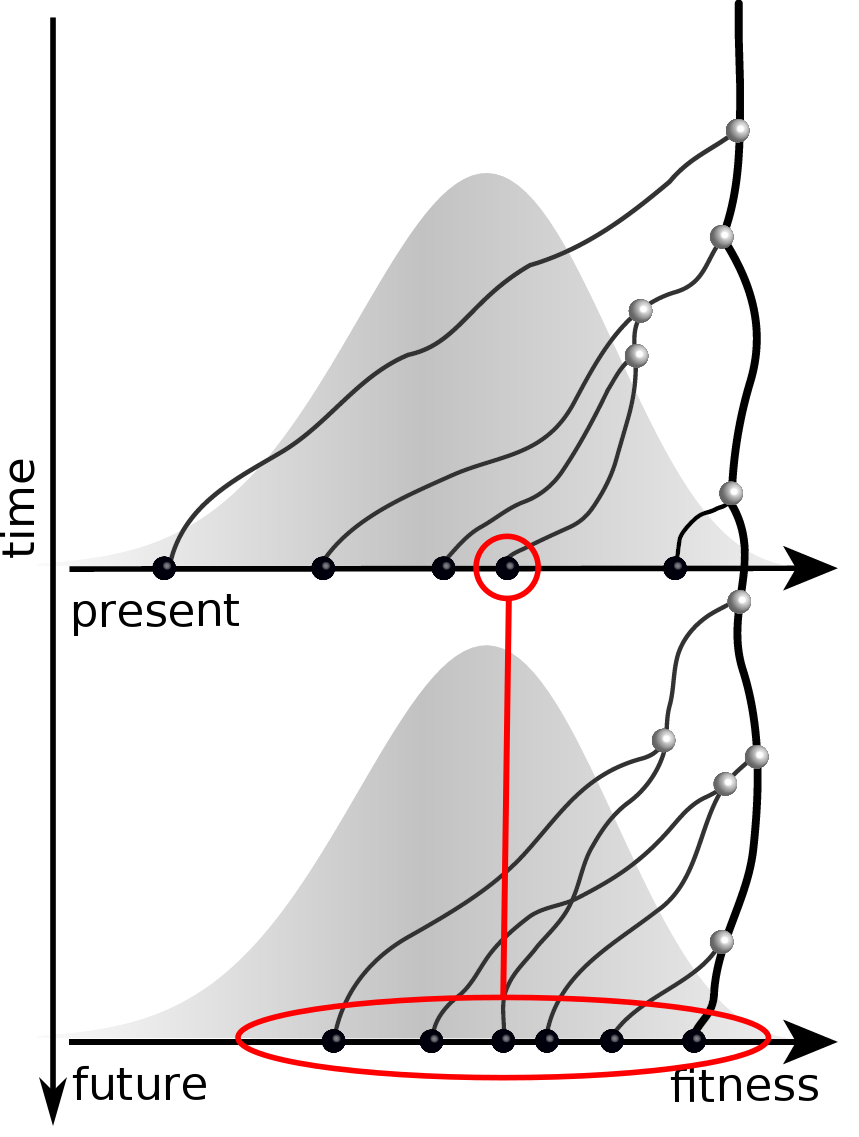
Bursts in a tree ↔ high fitness genotypes
Can we read fitness of a tree?



- Influenza virus evolves to avoid human immunity
- Vaccines need frequent updates

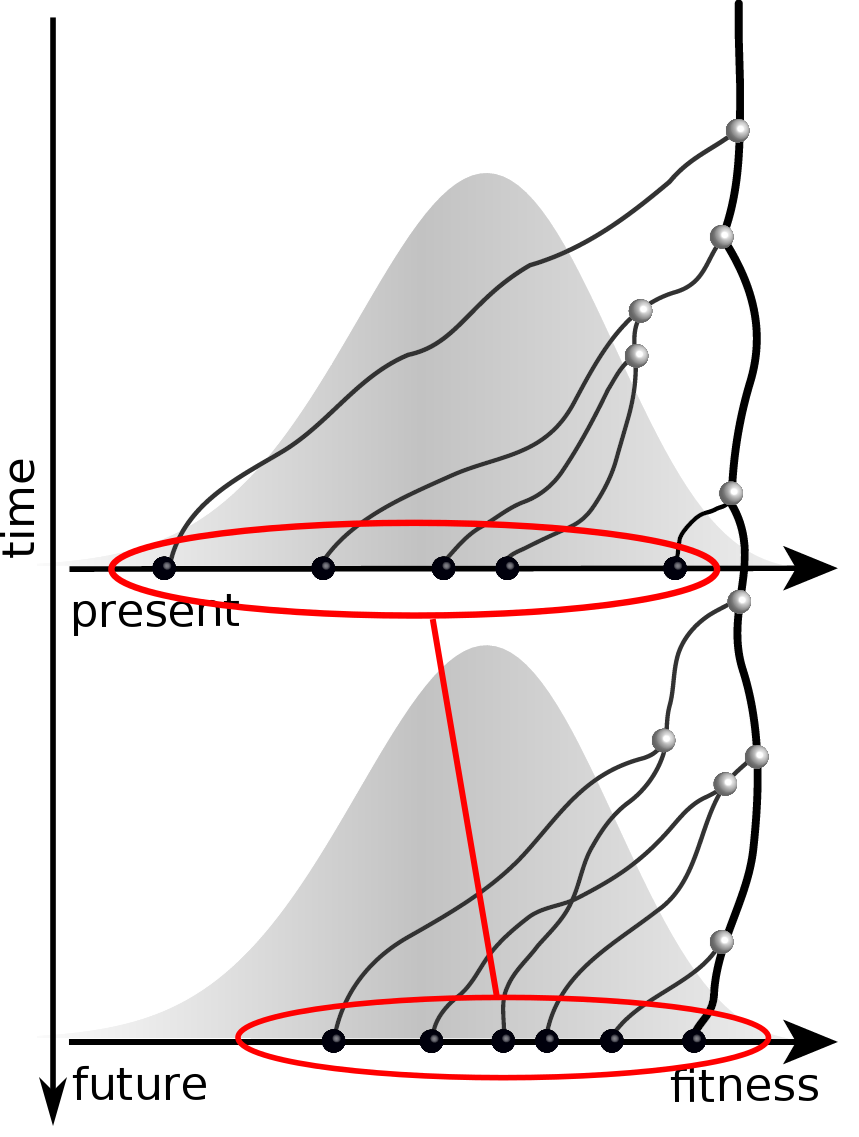
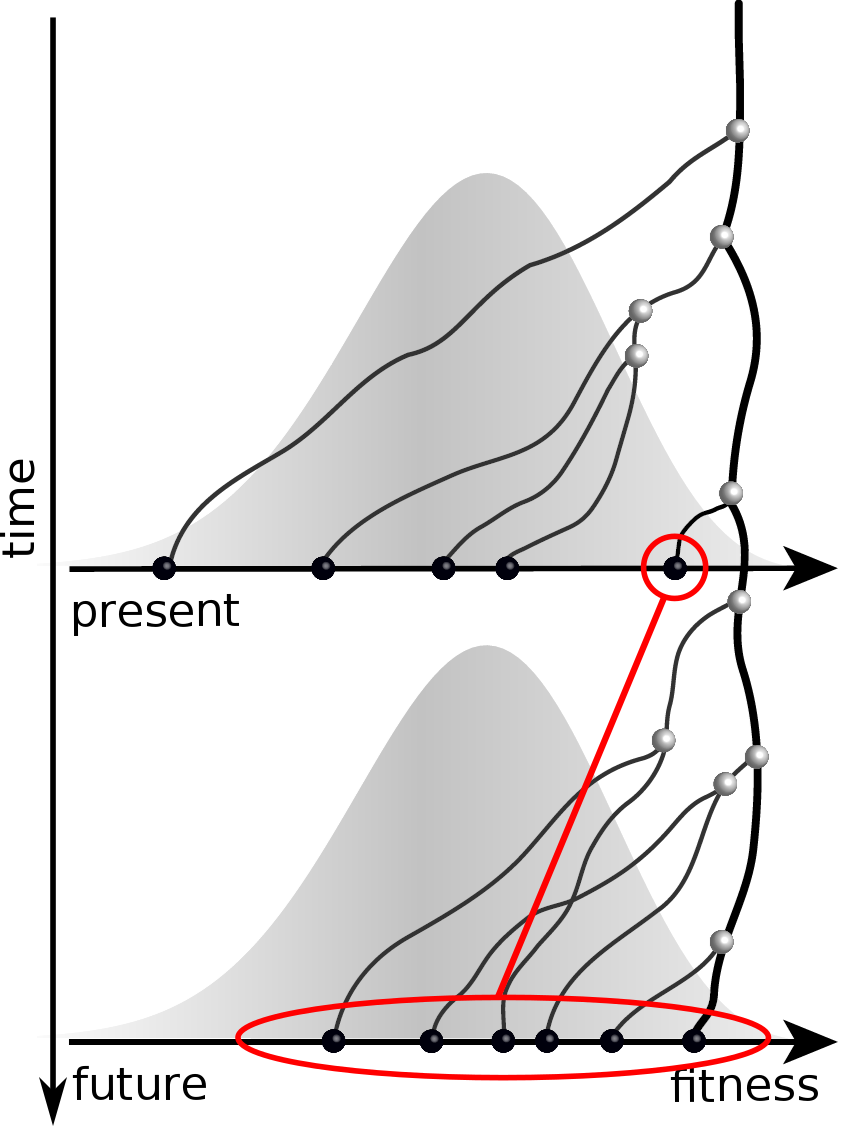
Predicting evolution

Given the branching pattern,
RN, Russell, Shraiman, eLife, 2014
- can we predict fitness?
- pick the closest relative of the future?
Fitness inference from trees
$$P(\mathbf{x}|T) = \frac{1}{Z(T)} p_0(x_0) \prod_{i=0}^{n_{int}} g(x_{i_1}, t_{i_1}| x_i, t_i)g(x_{i_2}, t_{i_2}| x_i, t_i)$$
RN, Russell, Shraiman, eLife, 2014
Validate on simulation data
- simulate evolution
- sample sequences
- reconstruct trees
- infer fitness
- predict ancestor of future
- compare to truth

Validation on simulated data
Validation on simulated data
Prediction of the dominating H3N2 influenza strain
RN, Russell, Shraiman, eLife, 2014HIV acknowledgments






- Fabio Zanini
- Jan Albert
- Johanna Brodin
- Christa Lanz
- Göran Bratt
- Lina Thebo


Influenza and Theory acknowledgments




- Boris Shraiman
- Colin Russell
- Trevor Bedford
- Oskar Hallatschek


My lab will move to the Biozentrum Basel
