Population genetics of rapid adaptation and fitness inference from trees
Richard Neher
Biozentrum, University of Basel
slides at neherlab.org/201905_phyloseminar.html
Dynamics, genetic diversity, and phylogenetic trees
evolutionary processes ↔ trees ↔ genetic diversity
Neutral models
- all individuals are identical → same offspring distribution
- Kingman coalesence emerges as universal description
- everything is easy to calculate
- generalization to structured coalescent
- generalization to multiple fitness classes possible
→ background selection
But: completely unsuitable to treat adaptive evolution!
Neutral/Kingman coalescent
strong selection
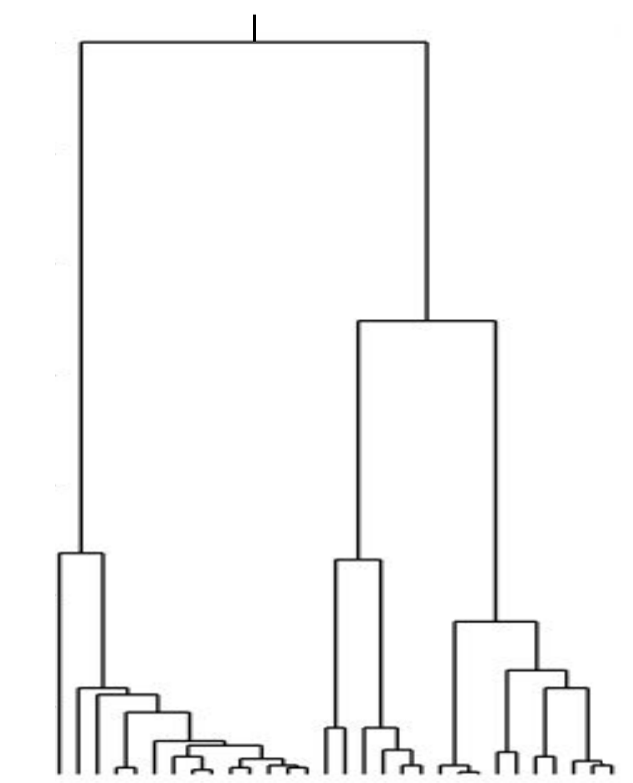
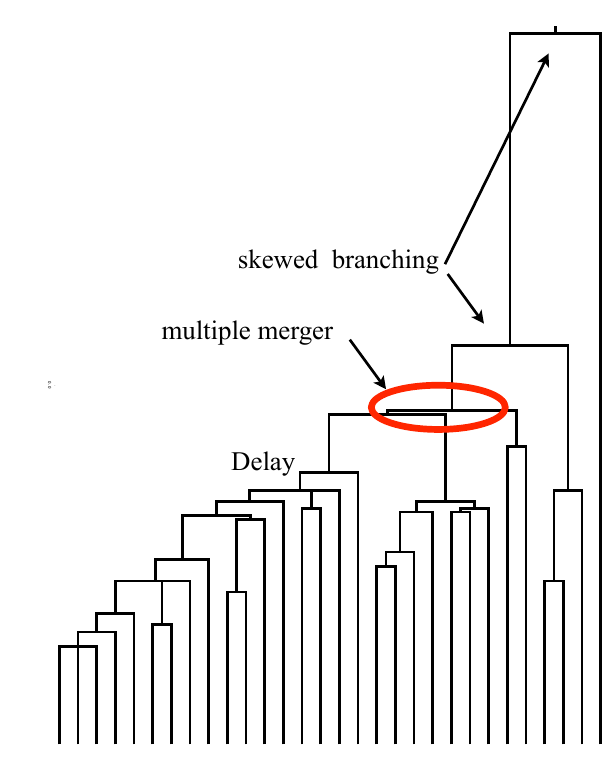
Fitness variation in rapidly adapting populations
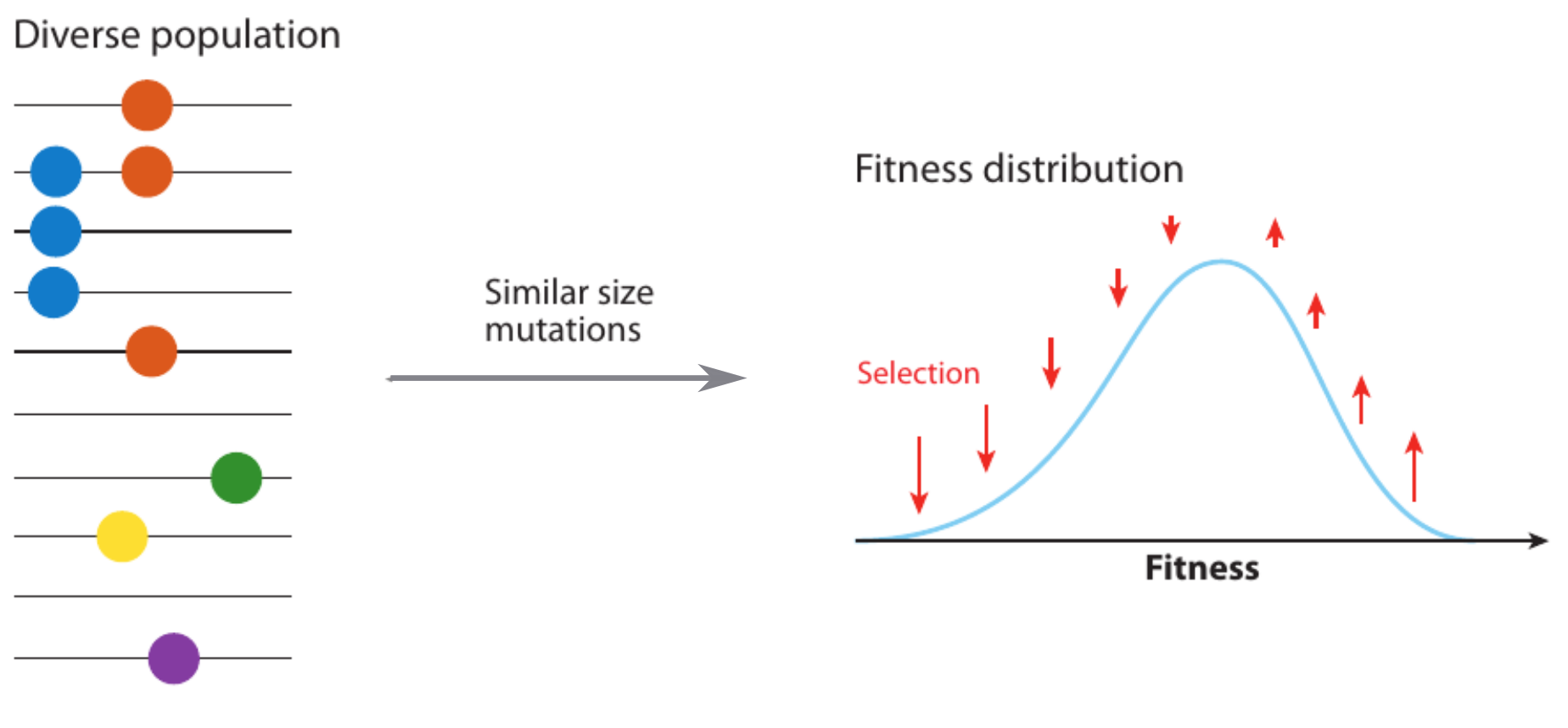
Traveling wave models of adaptation
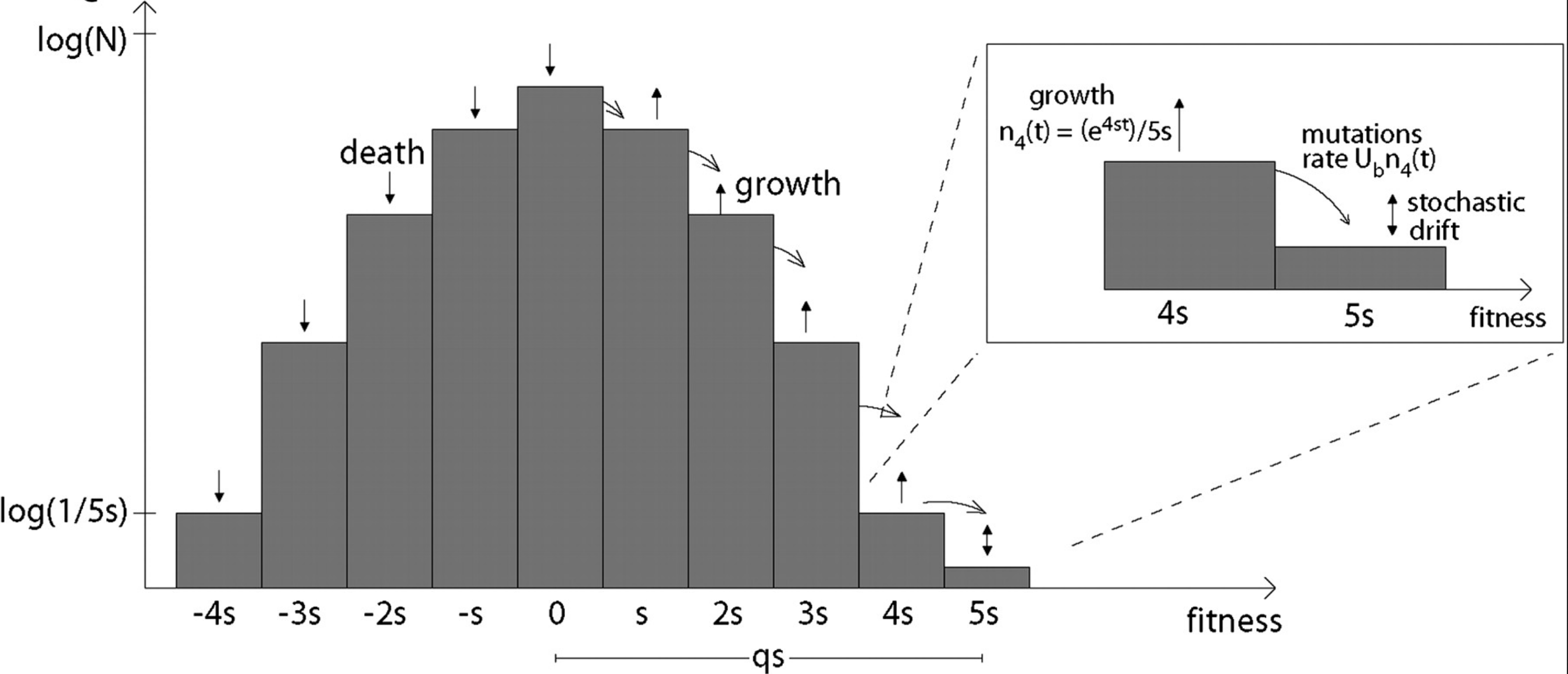
- Speed of adaptation is logarithmic in population size
- Environment (fitness landscape), not mutation supply, determines adaptation
- Different models have universal emerging properties

Bolthausen-Sznitman Coalescent
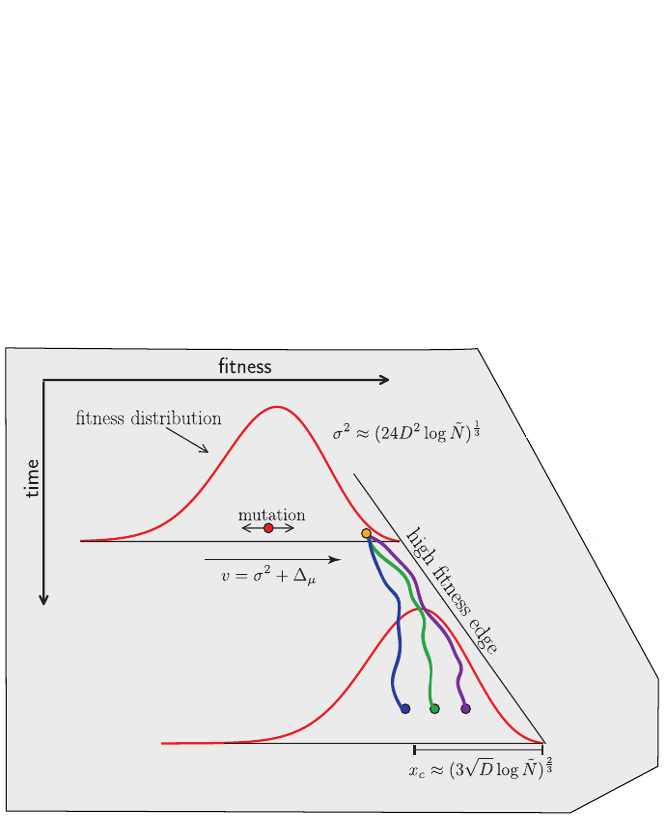
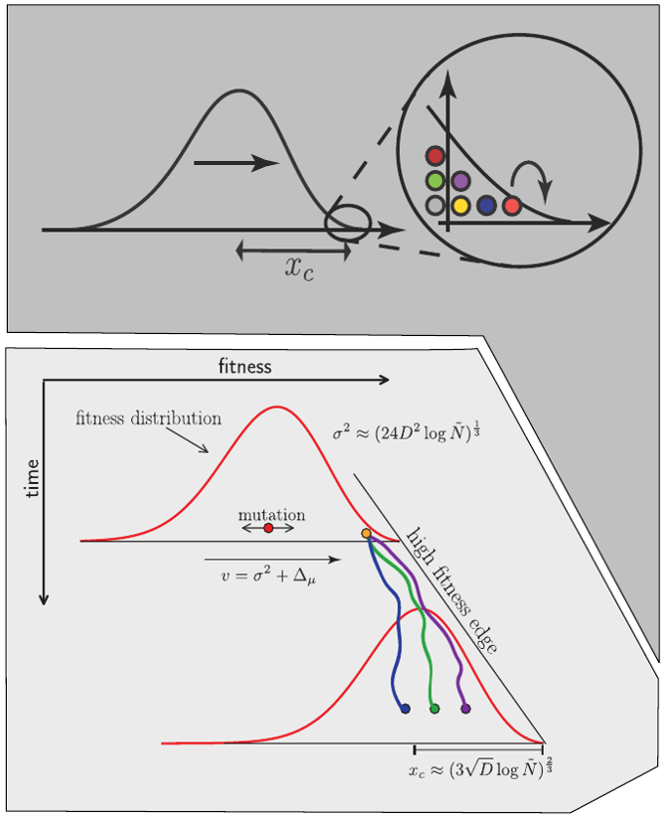
Traveling waves and the Bolthausen-Snitman coalescent
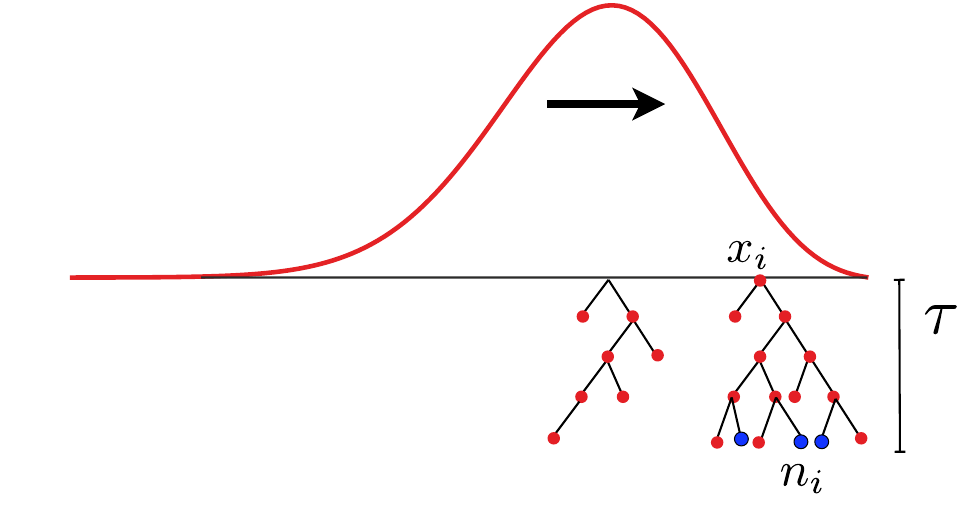
- Branching process approximation: $P(n_i, t|x_i)$
- Does a sample (blue dots) have a common ancestor $\tau$ generations ago? $\quad Q_b = \langle \sum_i \left(\frac{n_i}{\sum_j n_j}\right)^b\rangle \approx \frac{\tau-T_c}{T_c(b-1)} $
- Non-exchangeable at short times: fitness is inherited, lineages grow at different speed
- On intermediate time scales: exchangeable with power-law tail distribution: $P(n_i) \sim n_i^{-2}$
Universality of the Bolthausen-Sznitman Coalescent (BSC)
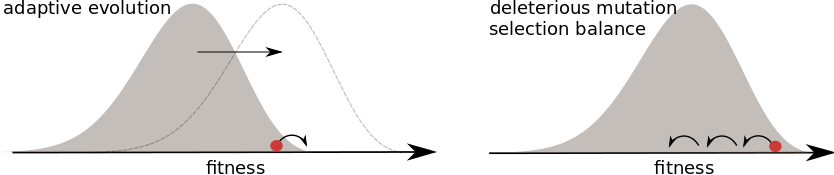
- many small effect mutations → coalescence is BSC like
- fitness diversity $\sigma$, not population size determines $T_{MRCA}$
- the time scale of coalescence is always $T_c \sim \sigma^{-1}\sqrt{\log N}$
- frequency dynamics is not diffusive, but has Levy-flight properties
- Can be extended to sexual populations
U-shaped polarized site frequency spectra
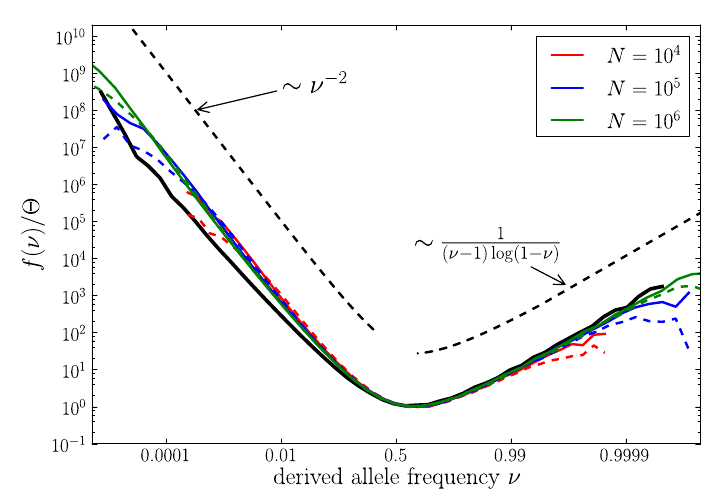

Universality -- adaptation and deleterious mutations
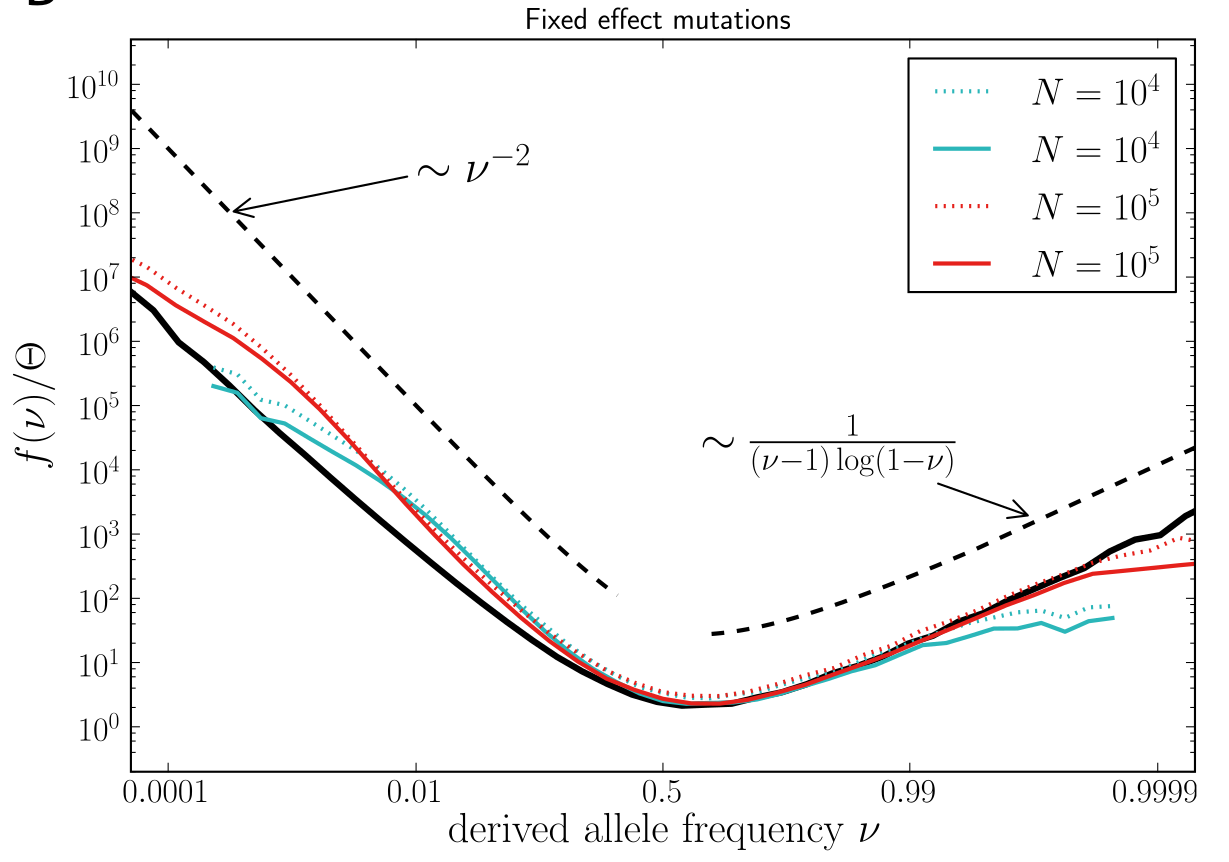
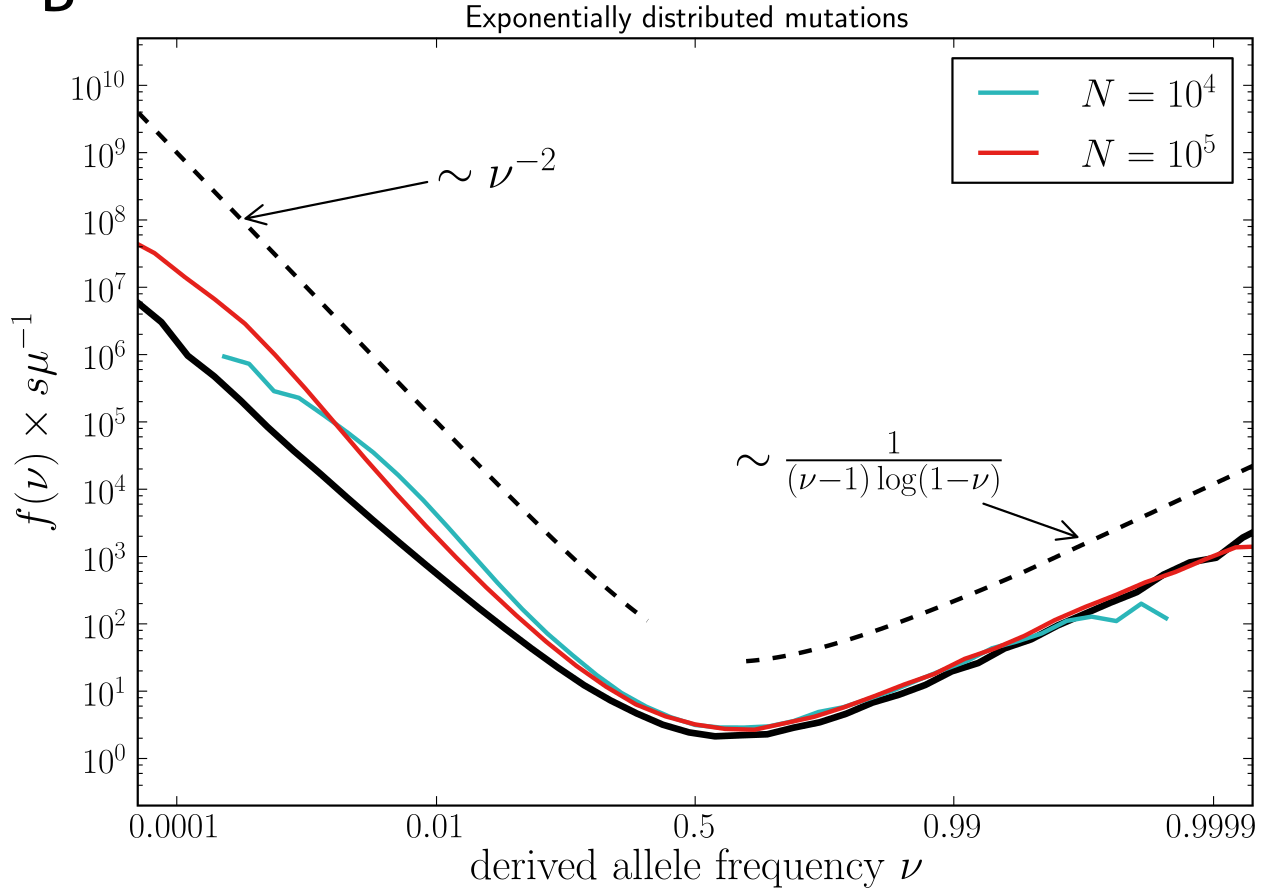
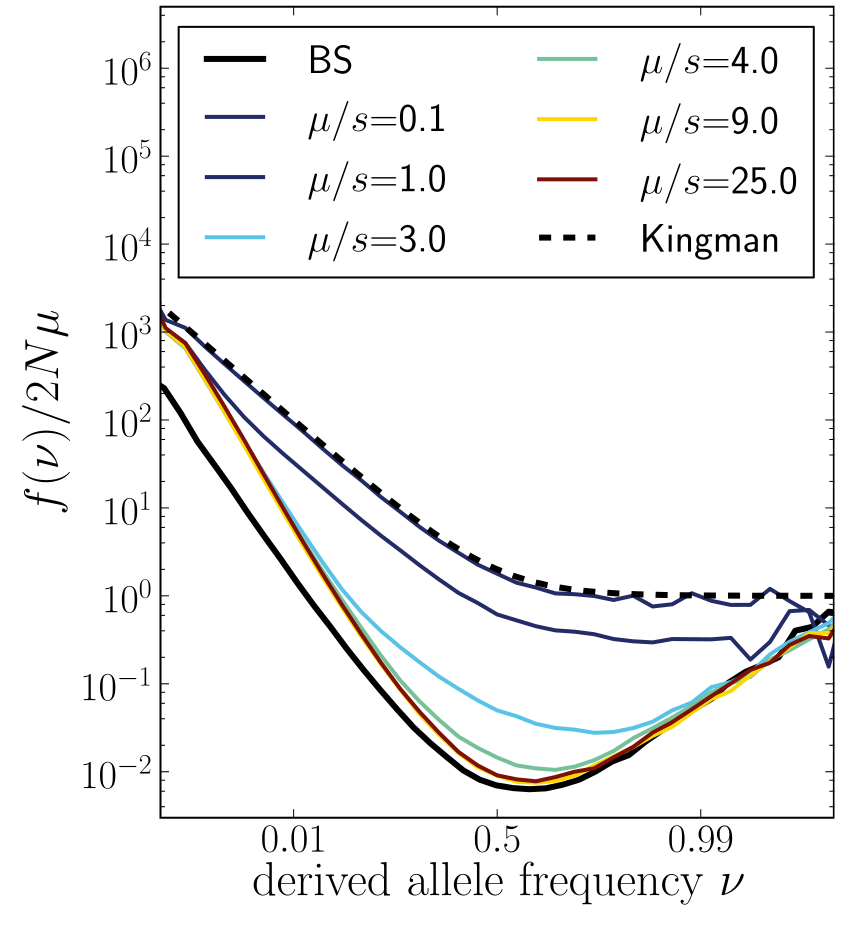
Bursts in a tree ↔ high fitness genotypes
Can we read fitness of a tree?
Predicting evolution

Given the branching pattern:
- can we predict fitness?
- pick the closest relative of the future?



- Influenza virus evolves to avoid human immunity
- Vaccines need frequent updates

Fitness inference from trees
$$P(\mathbf{x}|T) = \frac{1}{Z(T)} p_0(x_0) \prod_{i=0}^{n_{int}} g(x_{i_1}, t_{i_1}| x_i, t_i)g(x_{i_2}, t_{i_2}| x_i, t_i)$$
$g(x,t|y,t')$: density of observed child lineages at $(t, x)$ (time,fitness) given a parent at $(t',y)$
conditional no observed branching between parent and child.
RN, Russell, Shraiman, eLife, 2014
$g(x,t|y,t')$: density of observed child lineages at $(t, x)$ (time,fitness) given a parent at $(t',y)$
conditional no observed branching between parent and child.
Validate on simulation data
- simulate evolution
- sample sequences
- reconstruct trees
- infer fitness
- predict ancestor of future
- compare to truth

Validation on simulated data
Validation on simulated data
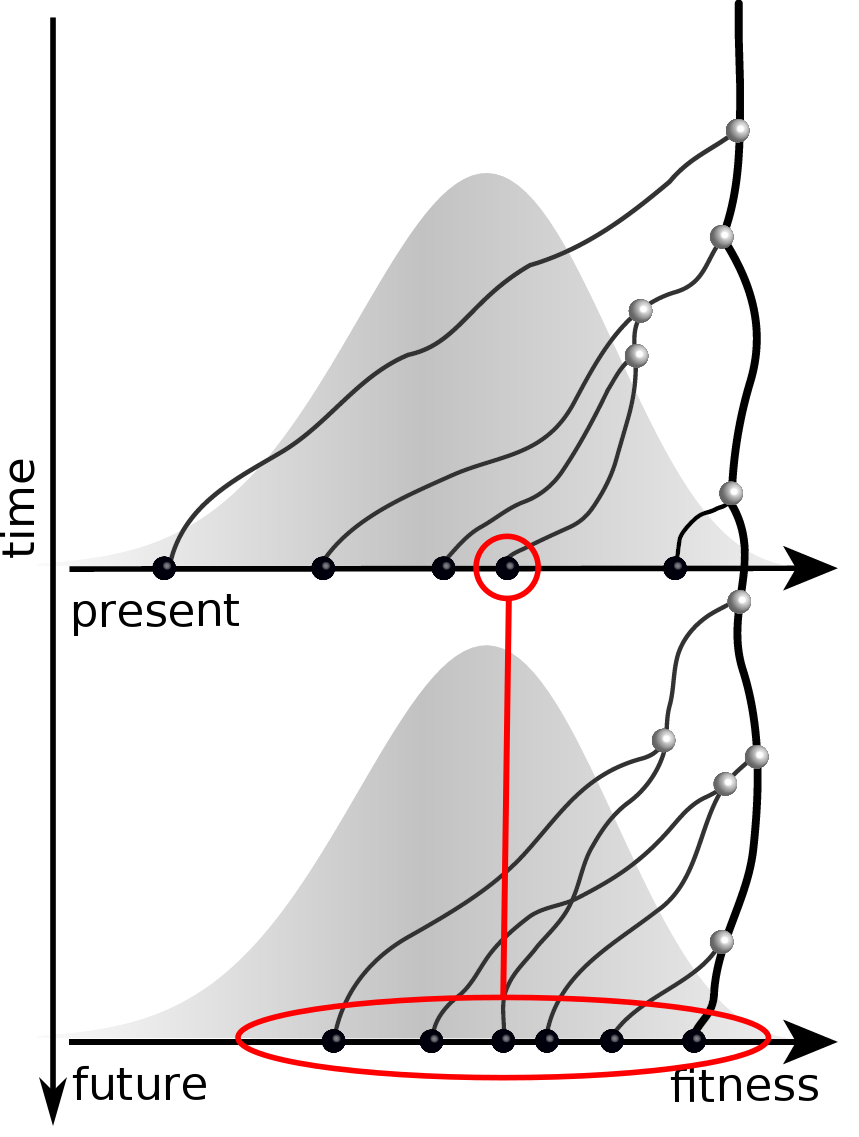
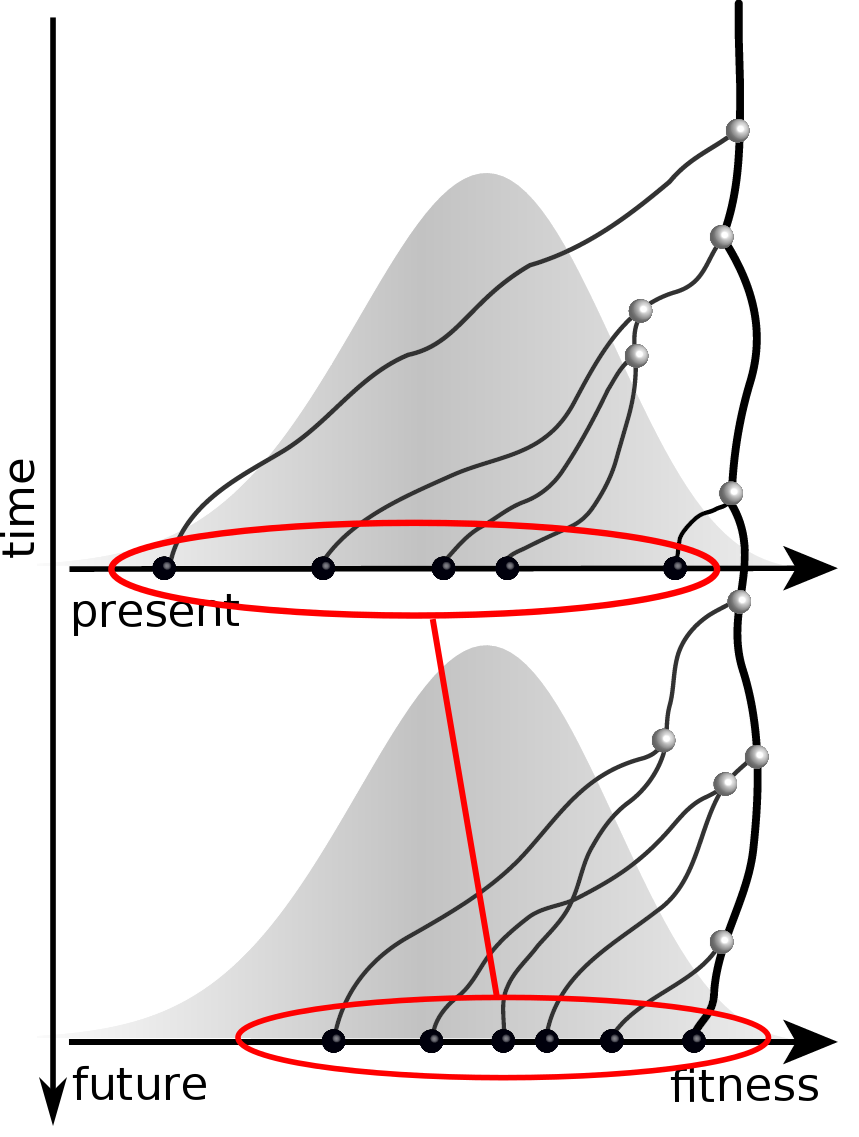
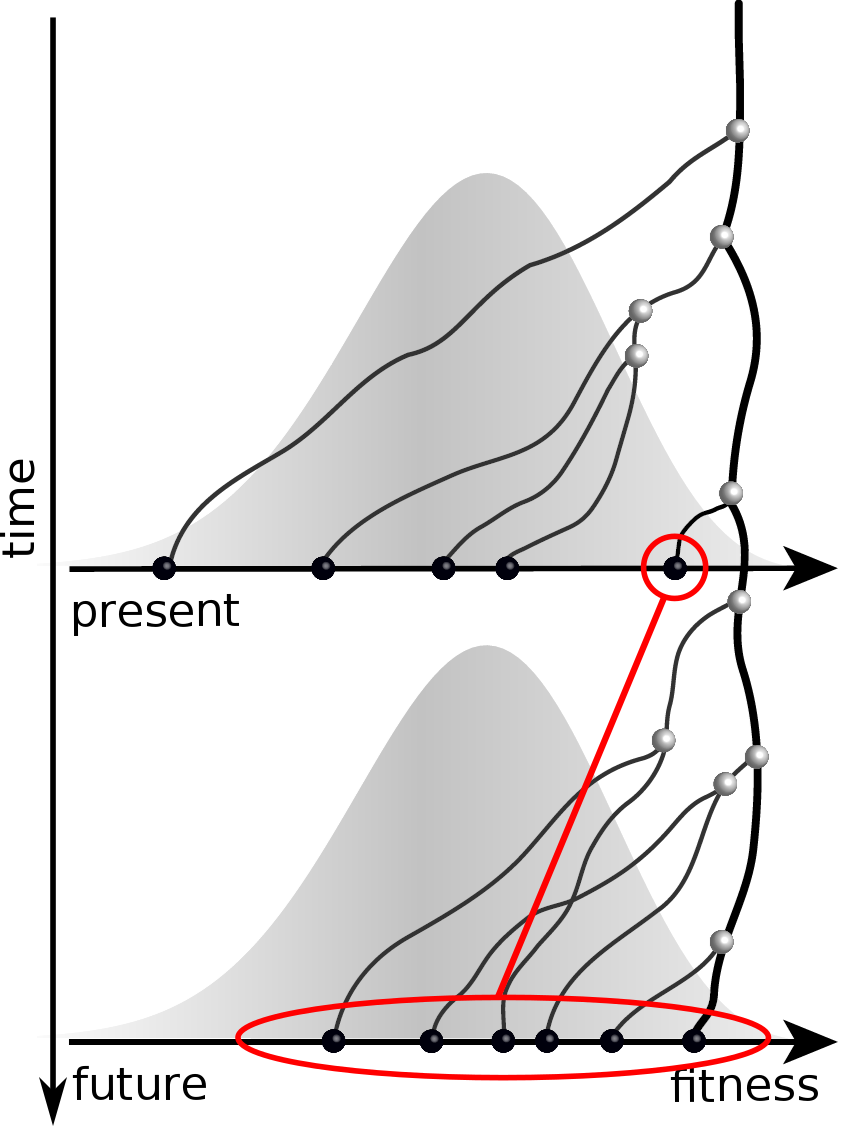
Prediction of the dominating H3N2 influenza strain
- no influenza specific input
- how can the model be improved? (see model by Luksza & Laessig)
- what other context might this apply?
Asymptotic analysis suggested simple heuristic: Local Branching Index
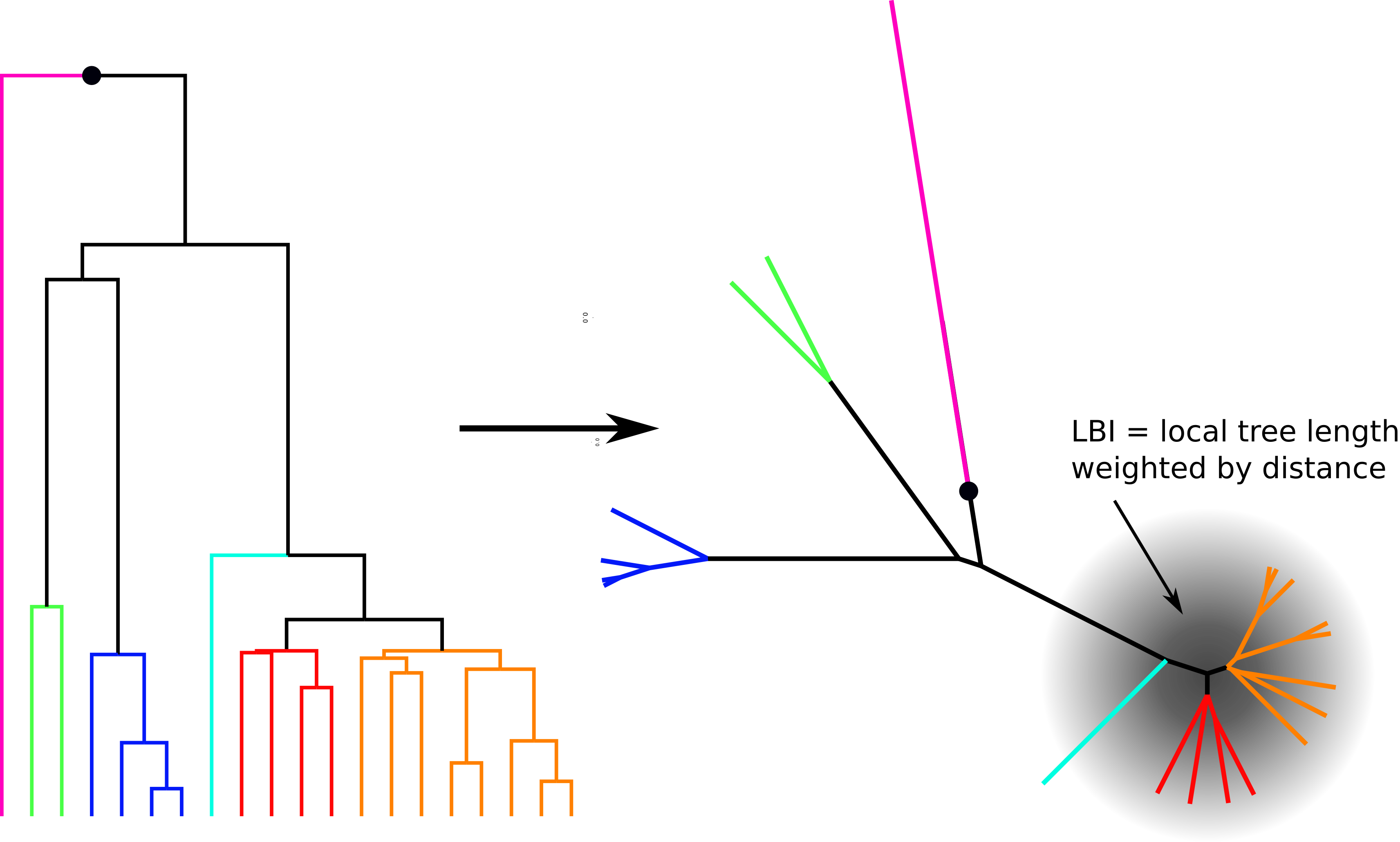

- For each node, calculate "tree volume" in neighborhood with an exponential kernel
- Characteristic scale: fraction of the coalescent time scale ($\sim T_c/15$)
Summary
- Fitness variation → no longer exchangeable pop-gen
- Mutational dynamics effectively restores exchangeability after some time, but....
- Resulting offspring distributions have long tails → no longer Kingman
- Bolthausen-Sznitman coalescent is universal if pioneer strains compete
- Insights into pop-gen of rapid adaptation → flu prediction

Acknowledgments



- Boris Shraiman (UCSB)
- Colin Russell (now Amsterdam)
- Oskar Hallatschek (UCB)



