Real-time tracking of RNA viruses using Nextstrain
Richard Neher
Biozentrum & SIB, University of Basel
slides at neherlab.org/202106_EPFL.html
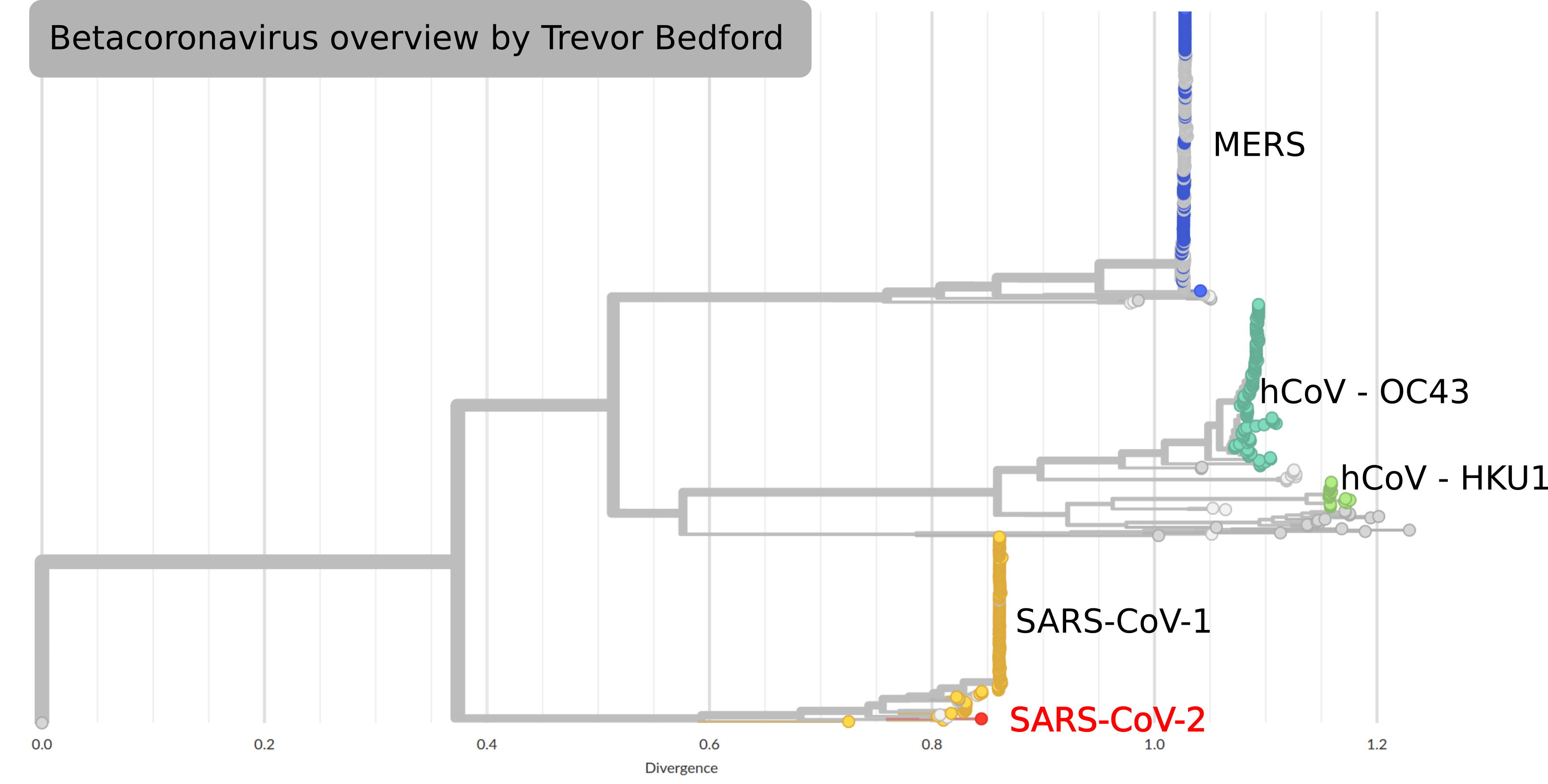 by Trevor Bedford
by Trevor Bedford
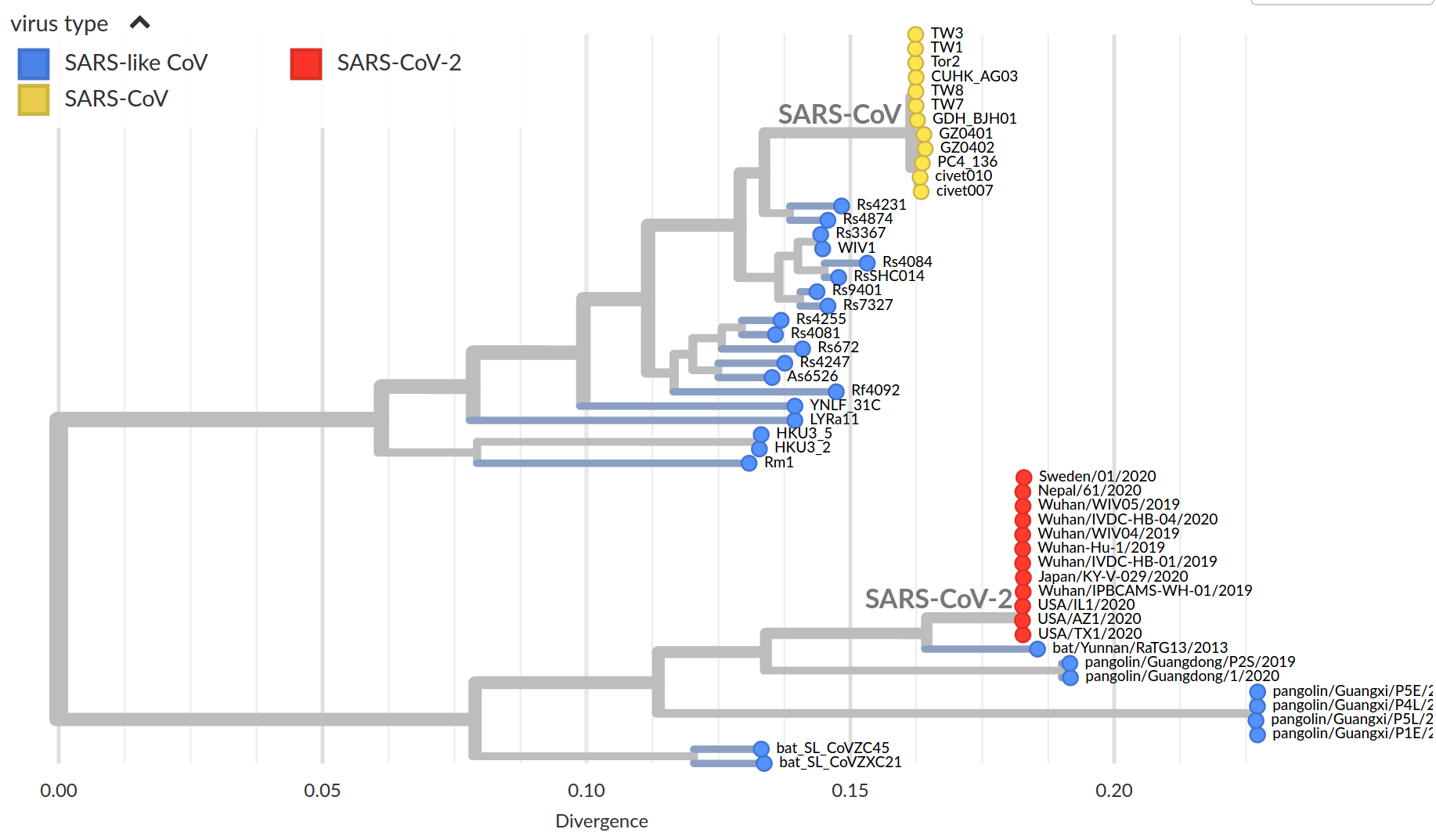 by Trevor Bedford
by Trevor Bedford
Tracking diversity and spread of SARS-CoV-2 in Nextstrain
Available data on Jan 26
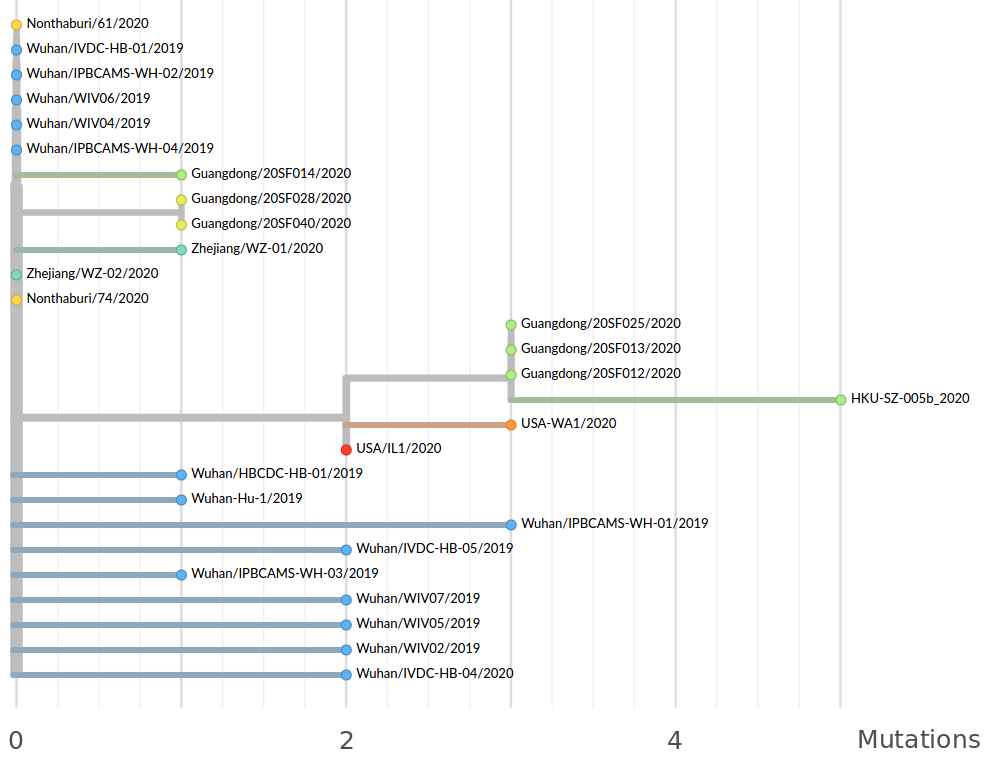
Early genomes differed by only a few mutations, suggesting very recent emergence
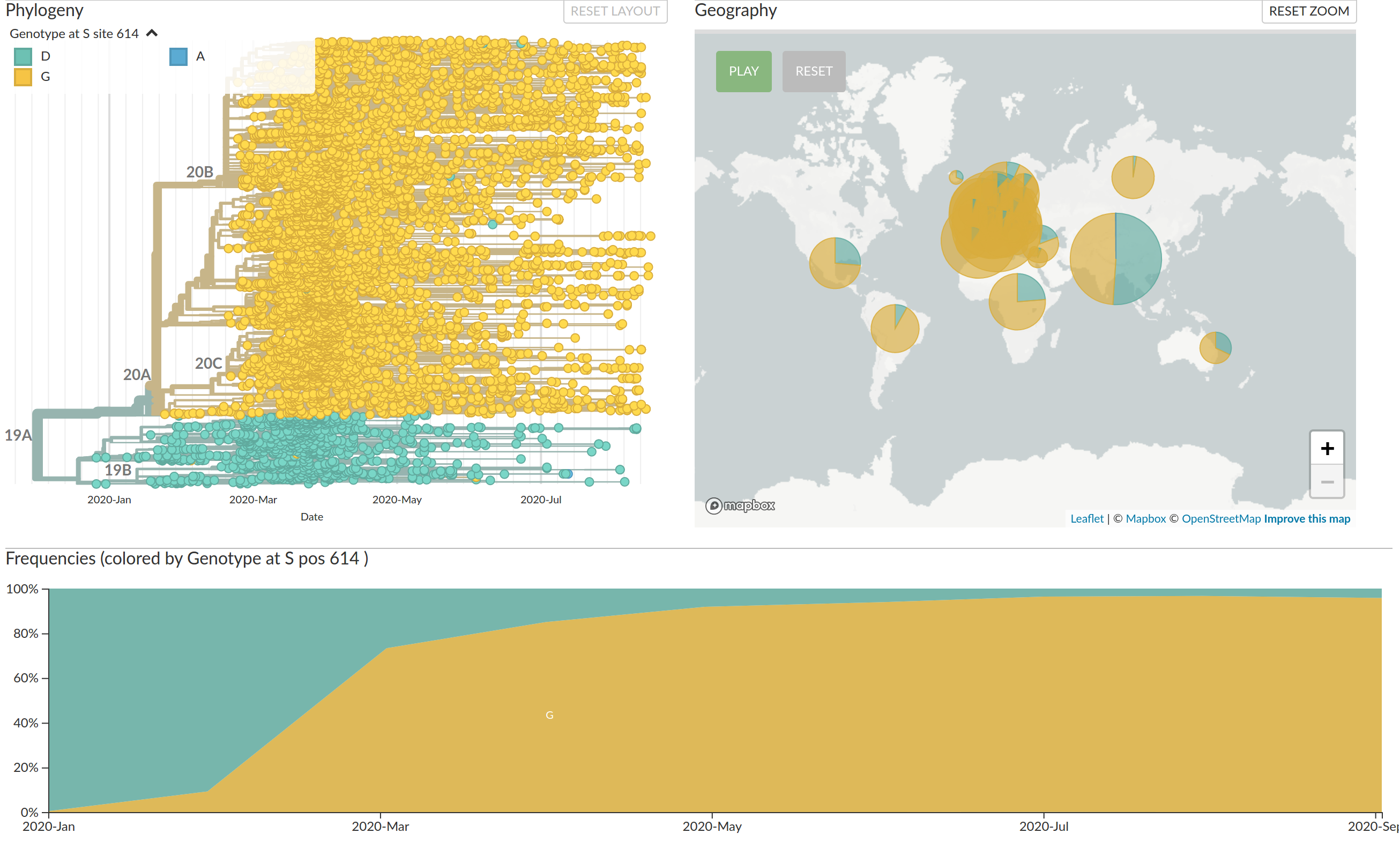
Interactive part on Nextstrain
VoCs have more mutations than expected...
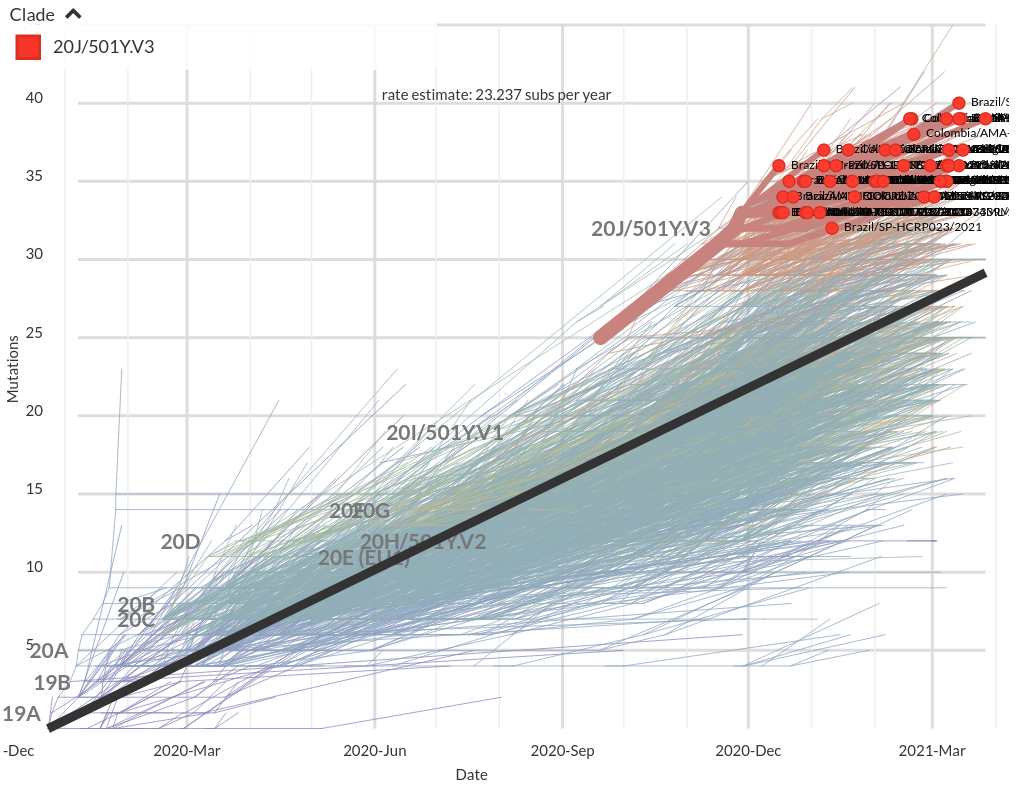 nextstrain
nextstrain
VoCs and VoIs: rapid converging evolution
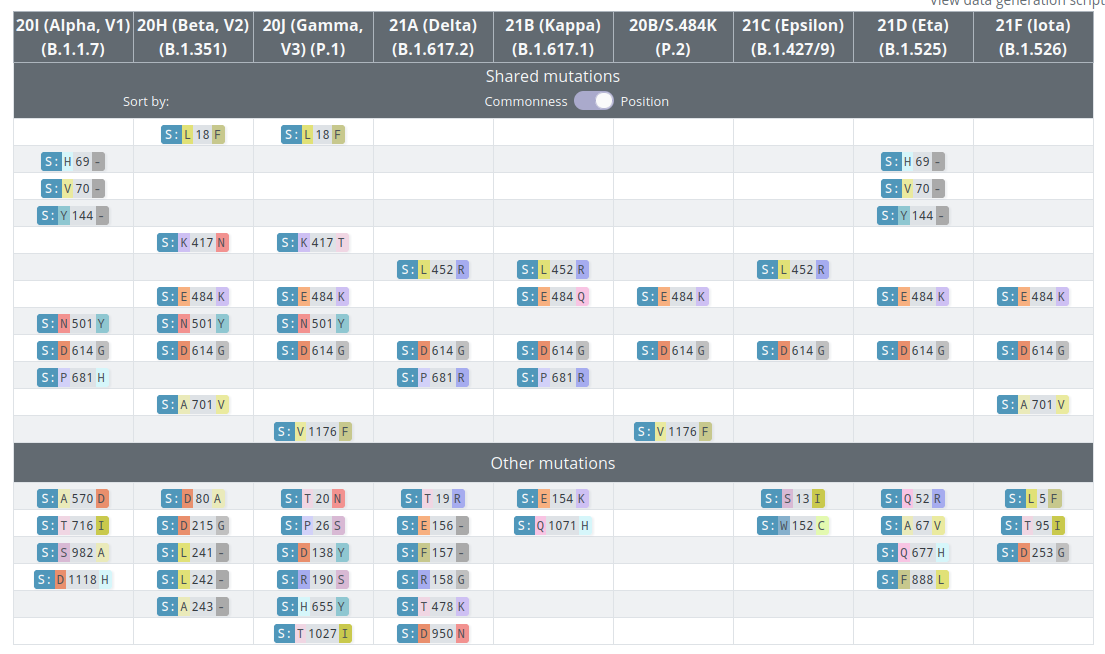 covariants.org by Emma Hodcroft
covariants.org by Emma Hodcroft
VoCs have been replacing other strains quickly
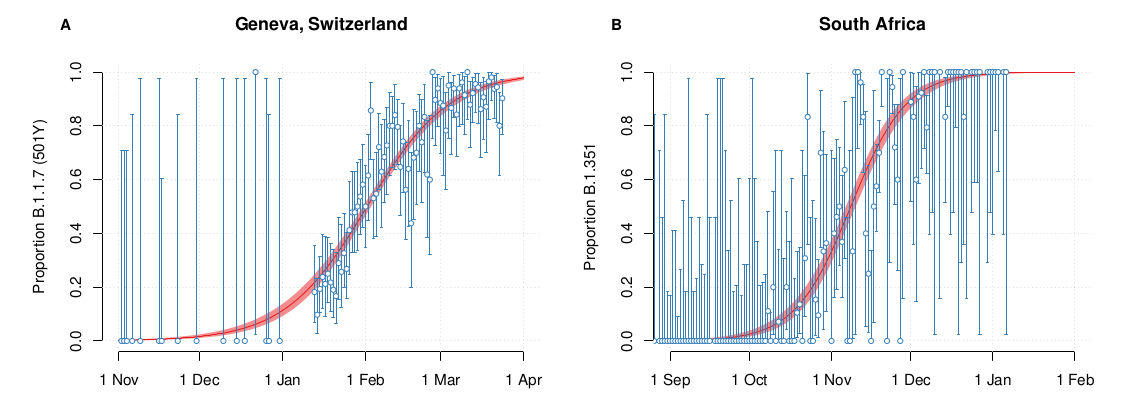
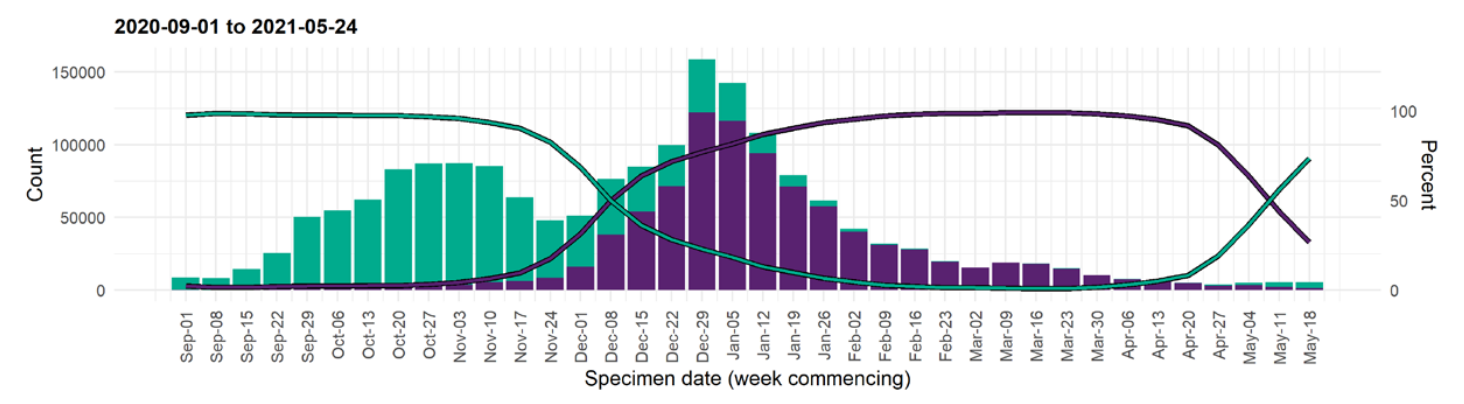 S-gene positivity in the UK
S-gene positivity in the UK
- How unusual is this?
- How do we tell early on whether a variant is more transmissible?
- How important are travel and seasons?
Recurrent mutations are common
- RNA viruses are not mutation limited
- In suitable environments, homoplasies are common
- Clearly selected variation, but they don't necessarily sweep
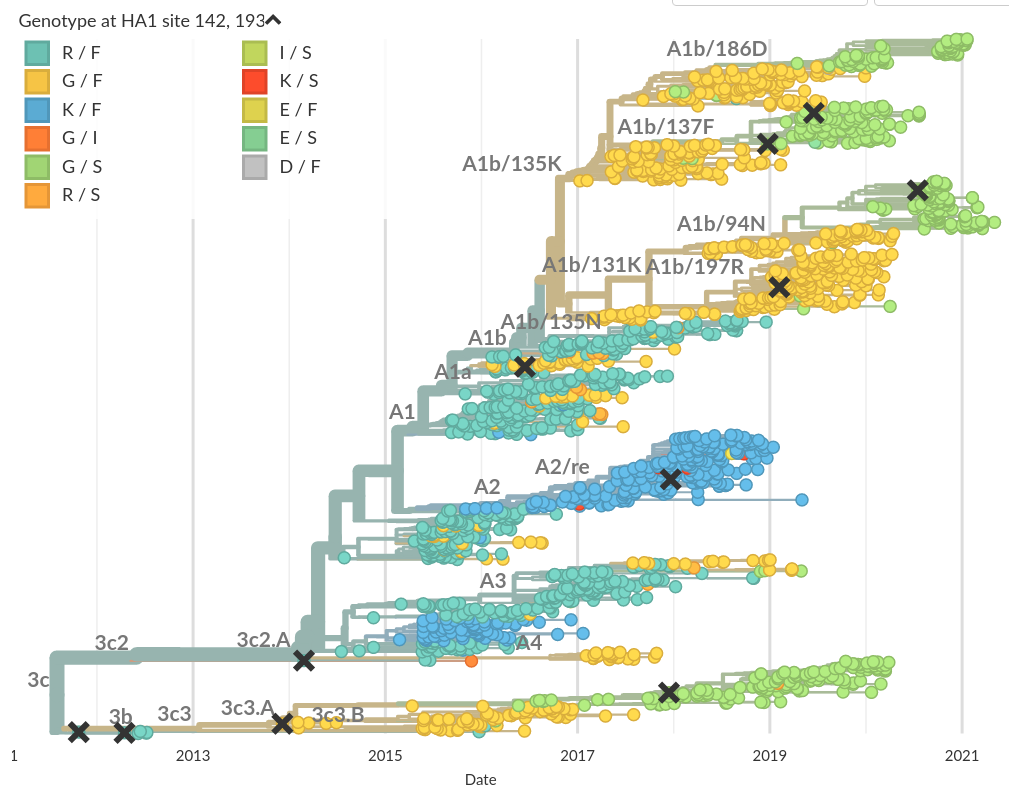
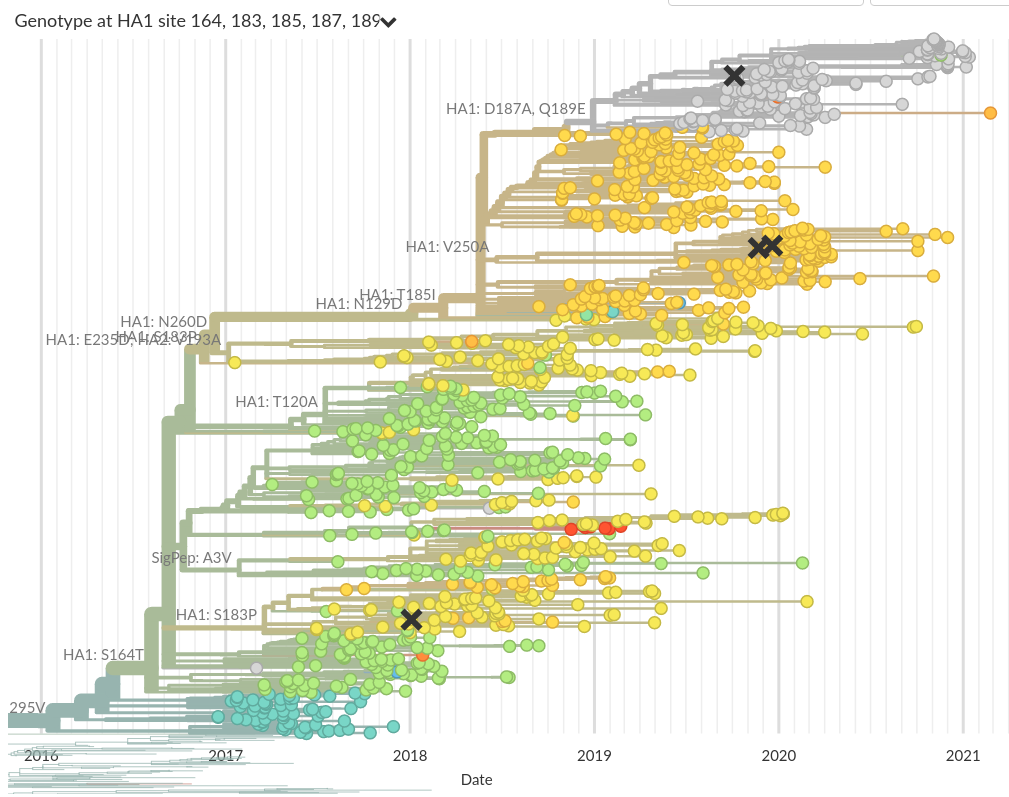
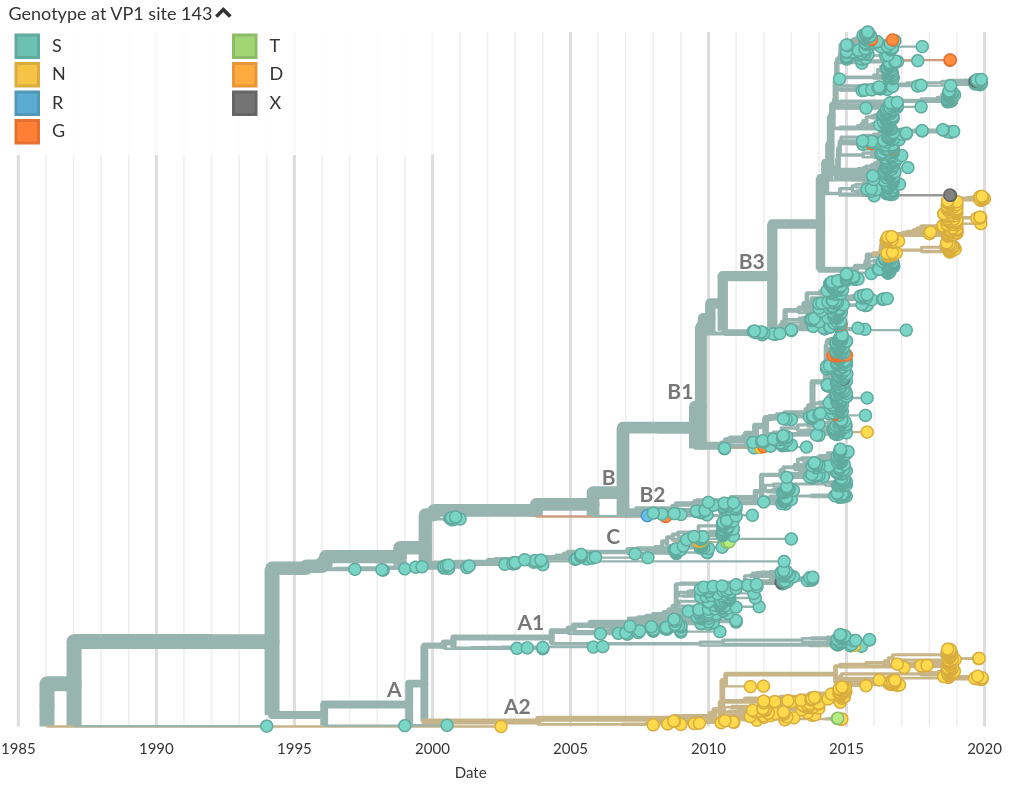
Do A/H3N2 mutations have inertia?
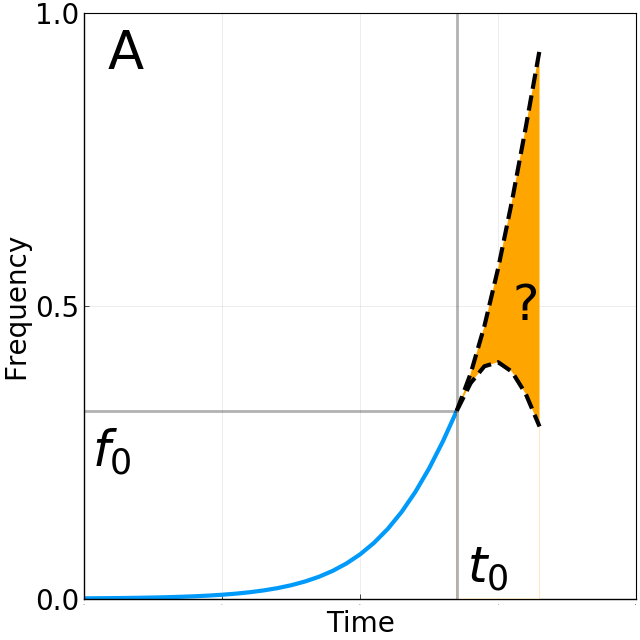
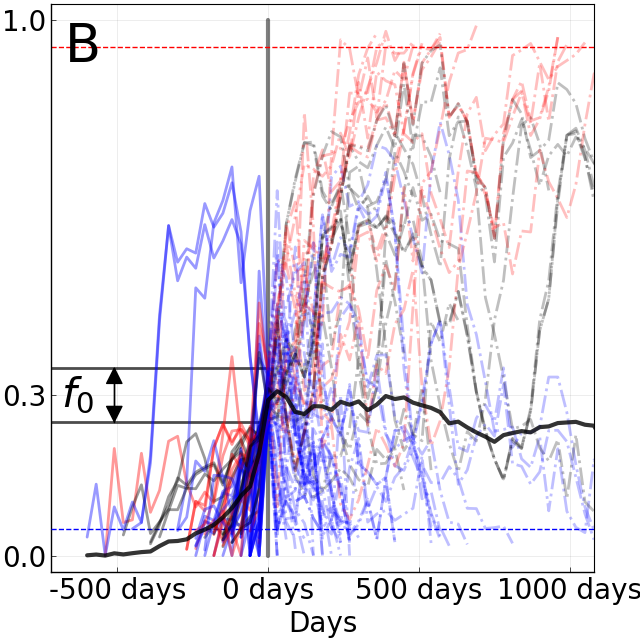
Fixation probability is quasi-neutral
A/H3N2 influenza
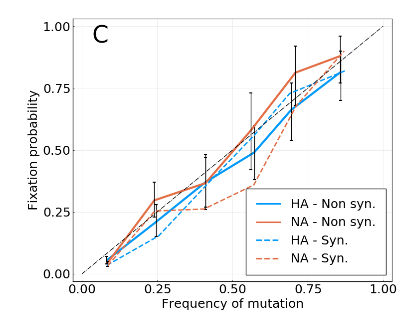
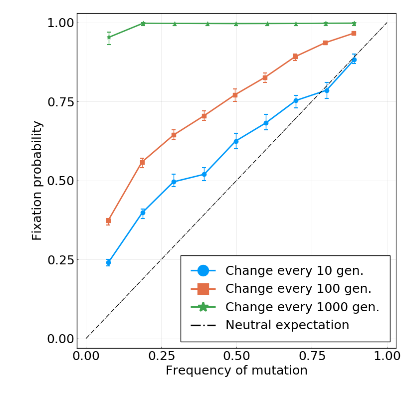
Simulations with increasing interference
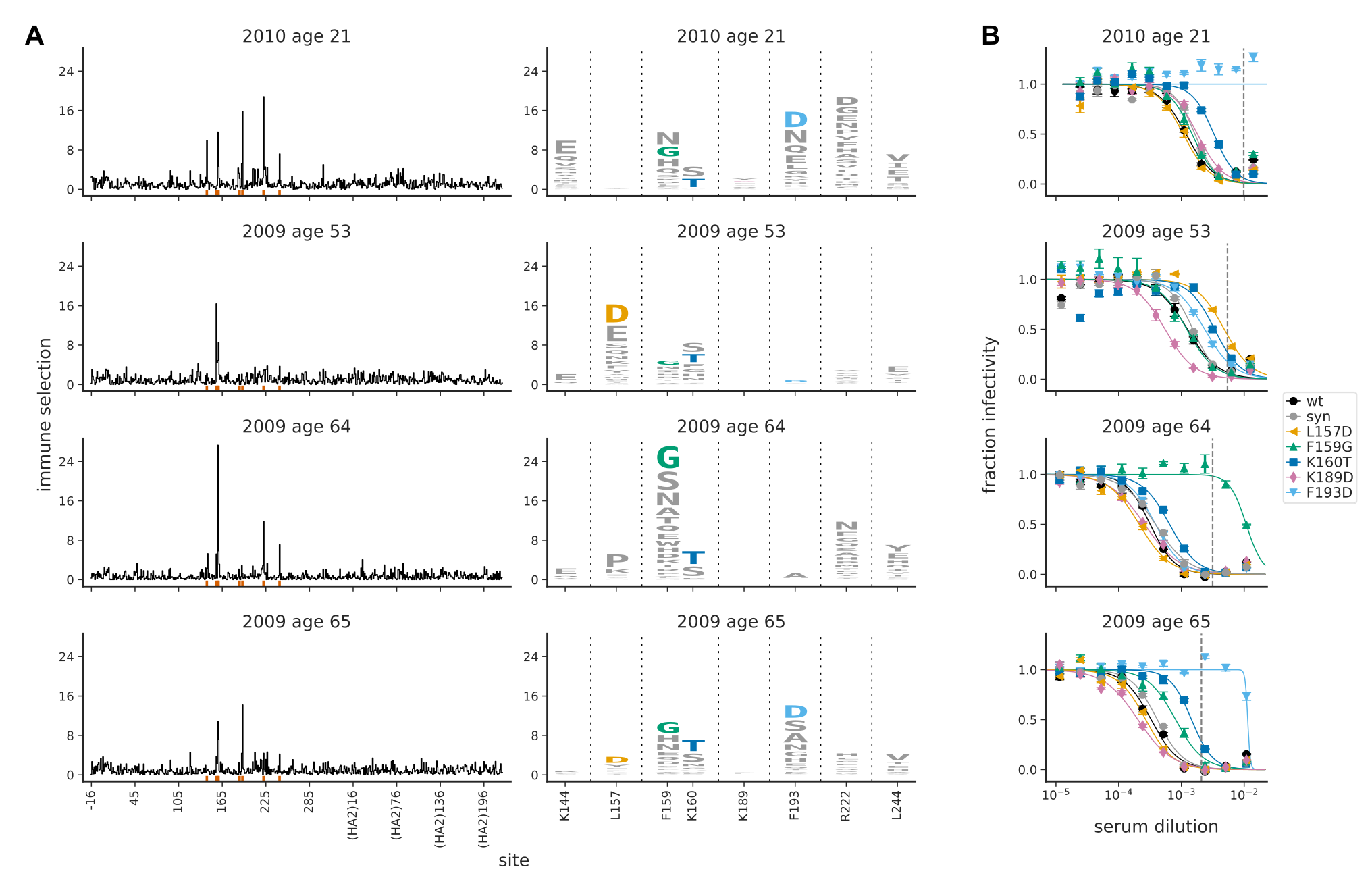
Diversity in immune responses
A/H3N2 influenza doesn't quite fit the textbook picture of rapid adaptation
- Strong signal of positive selection
- Evolutionary rate at antigenic sites very high
- Rapid clade displacement, interference
- Any given escape only allows to escape a fraction of the population
→ expiration of selection long before a mutation is common - The more diverse the immune landscape, the more neutral it looks
SARS-CoV-2 variants
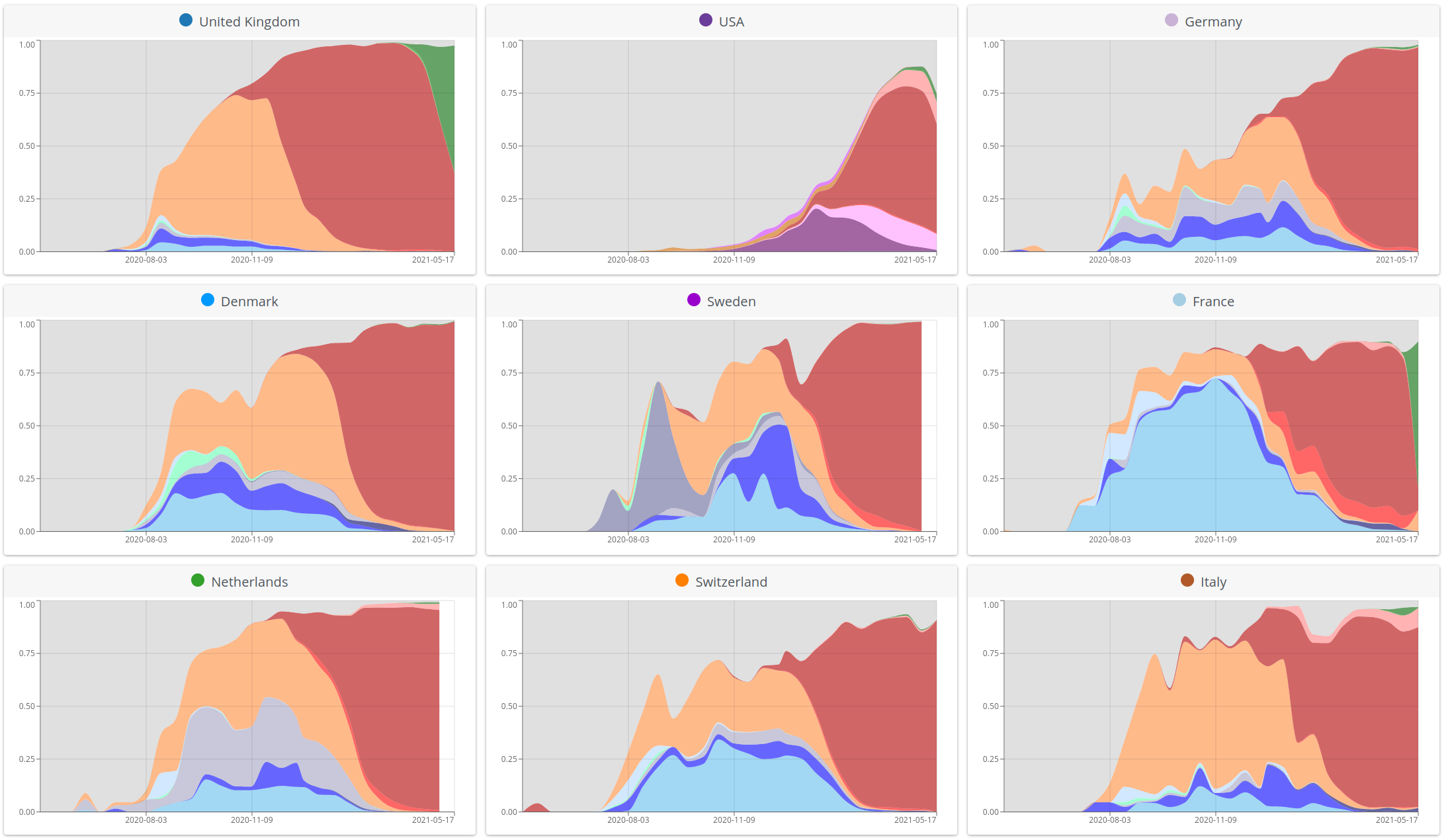
SARS-CoV-2 variants can become dominant without advantage
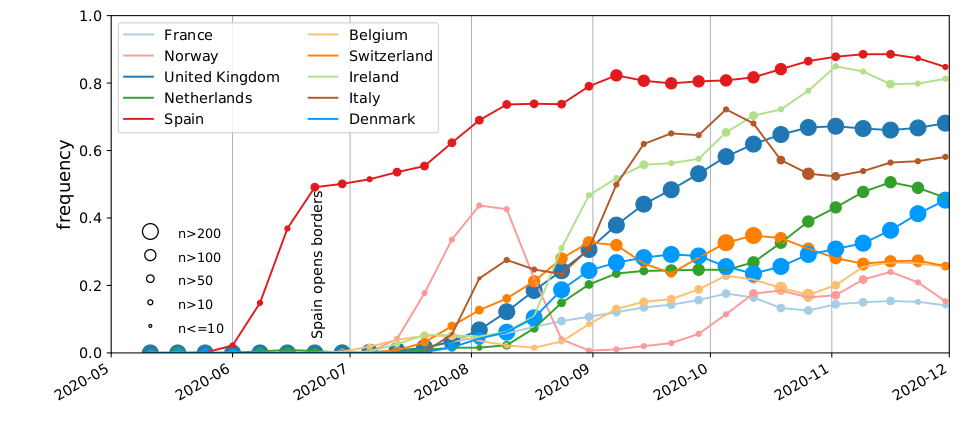
High case numbers in Spain and high travel volume spread the variant
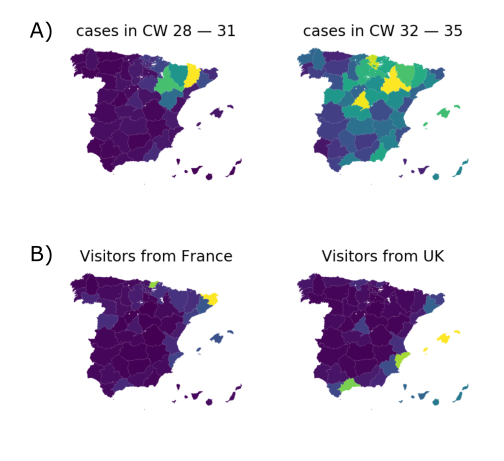
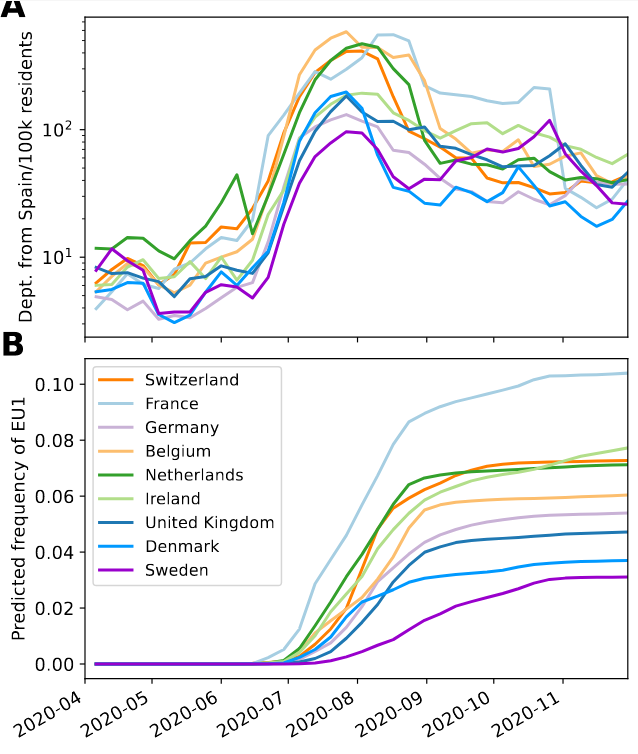
Travel driven variant dispersal and displacement
- High invidence differential and high travel volume can shift variant distribution
- Travel associated activities and behavior further increases impact
- Onward spread in traveling demographics can be higher
- We will see whether this also holds for 21A (delta)
- Very easy to jump to premature conclusions
Travel driven variant dispersal and displacement
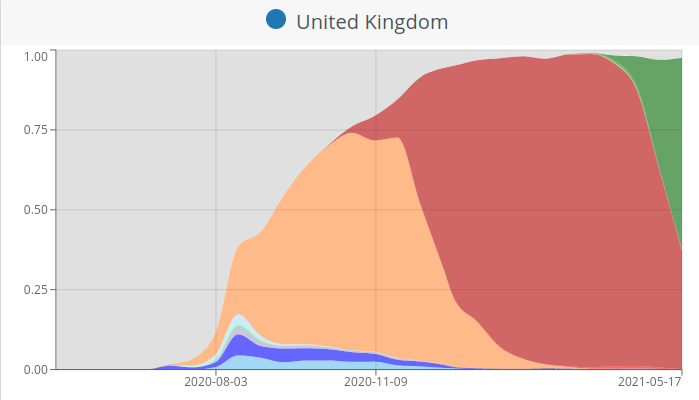
- rapid rise to dominance
- BUT: India had very high incidence
- travel between UK-India was high in March/April (put on red list on April 23rd)
- Likely hundreds of imports
- The variant is diverse and dates back to last summer.
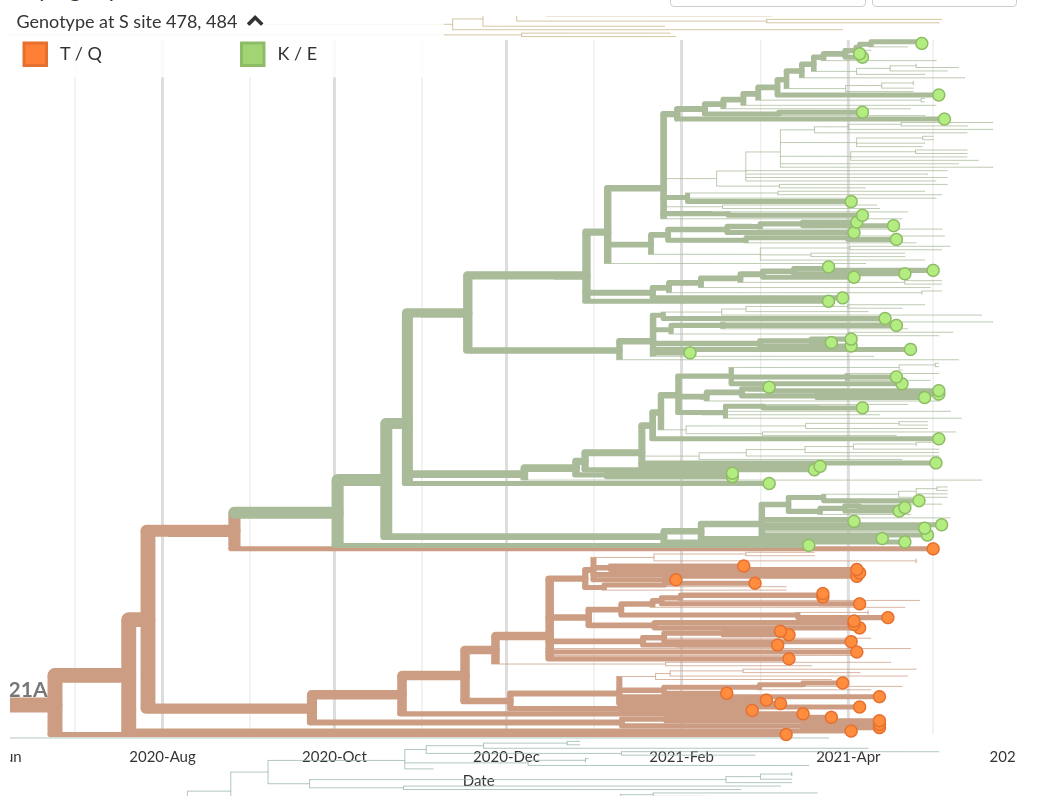
Transmission advantage relative to original variants likely.
But unclear how much.
Human corona viruses have pronounced seasonal prevalence (Sweden)
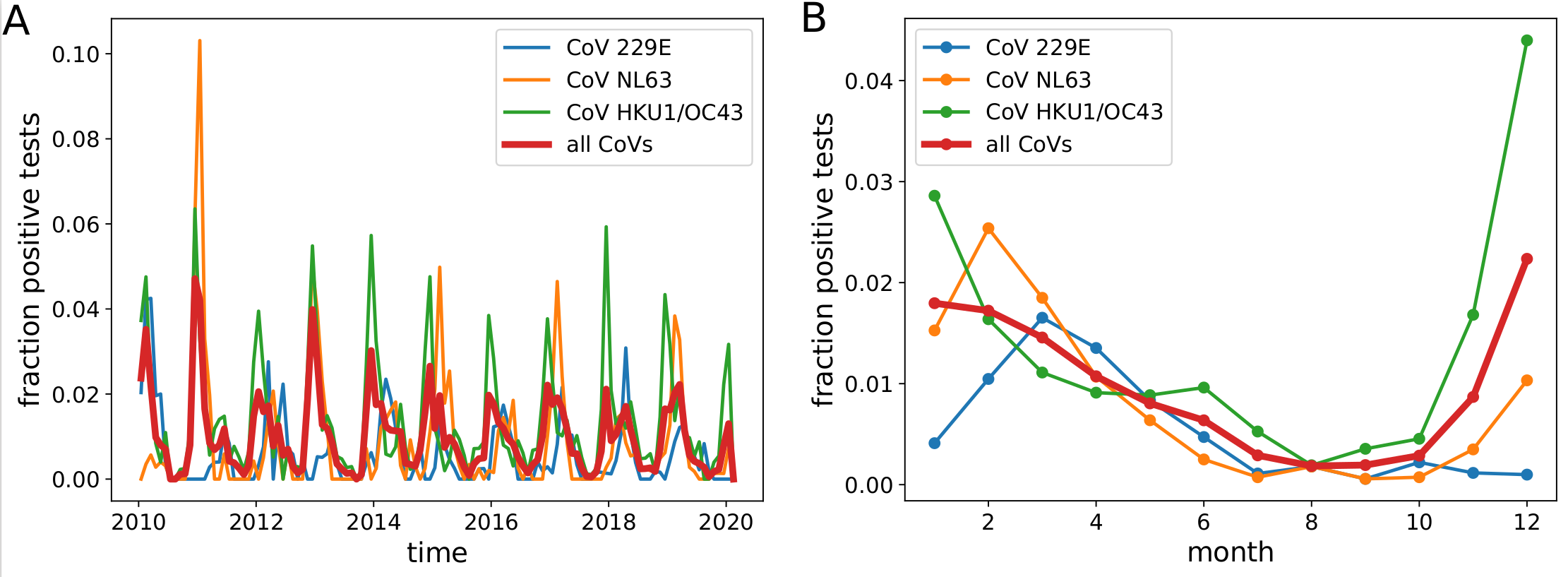
- Respiratory virus incl seasonal CoVs show seasonality
- Forcing through human behavior (indoor/outdoor activities).
- Control of the virus might be harder in temperate winter
- Absolute effect of seasonality unknown
Potential transition to an endemic seasonal virus (model from Feb 2020)
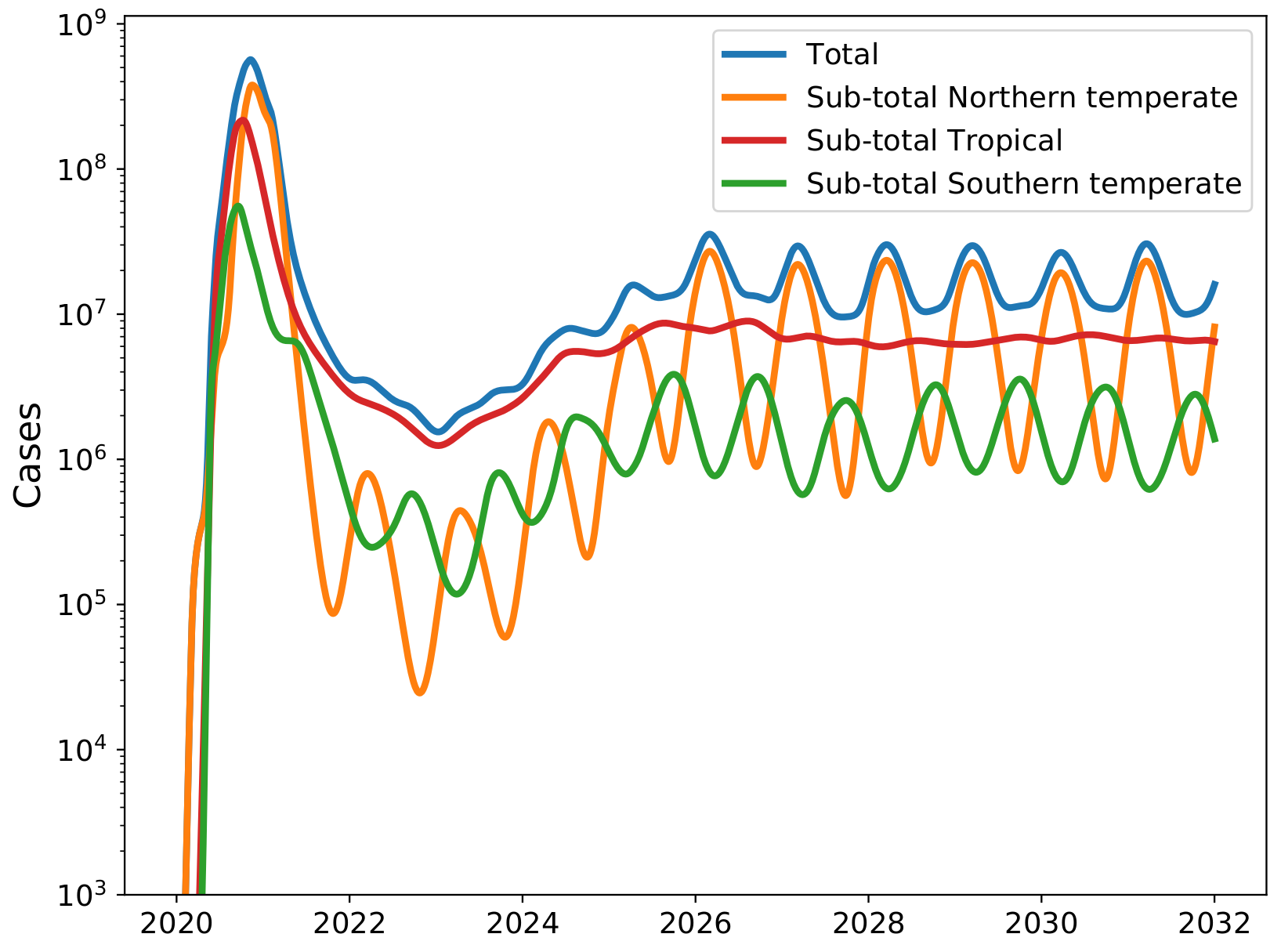
Influenza acknowledgements



- Boris Shraiman
- Trevor Bedford
- Pierre Barrat
- All the NICs and WHO CCs that provide influenza sequence data
- The WHO CCs in London and Atlanta for providing titer data


SARS-CoV-2 acknowledgements

.jpg)
- Emma Hodcroft (now in Bern)
- Moira Zuber (Basel)
- Iñaki Comas and Fernando Gonzalez-Candelas, Valencia
- Martina Reichmuth and Christian Althaus (Bern)
- Tanja Stadler, Sarah Nadeau, Tim Vaughan at ETH
- Alberto Hernando and David Matteo at Kido Dynamics
- Jesse Bloom, Katherine Crawford at Fred Hutch
- David Veesler, Alex Walls, Davide Corti, John Bowen at UW

Acknowledgments
Trevor Bedford and his lab -- terrific collaboration since 2014

especially James Hadfield, Emma Hodcroft, Ivan Aksamentov, Moira Zuber, John Huddleston, and Tom Sibley
Data we analyze are contributed by scientists from all over the world
Data are shared and curated by GISAID

