Tracking and predicting SARS-CoV-2 variants with Nextstrain
Richard Neher
Biozentrum & SIB, University of Basel
slides at neherlab.org/202107_CSHL.html
Available data on Jan 26 2020
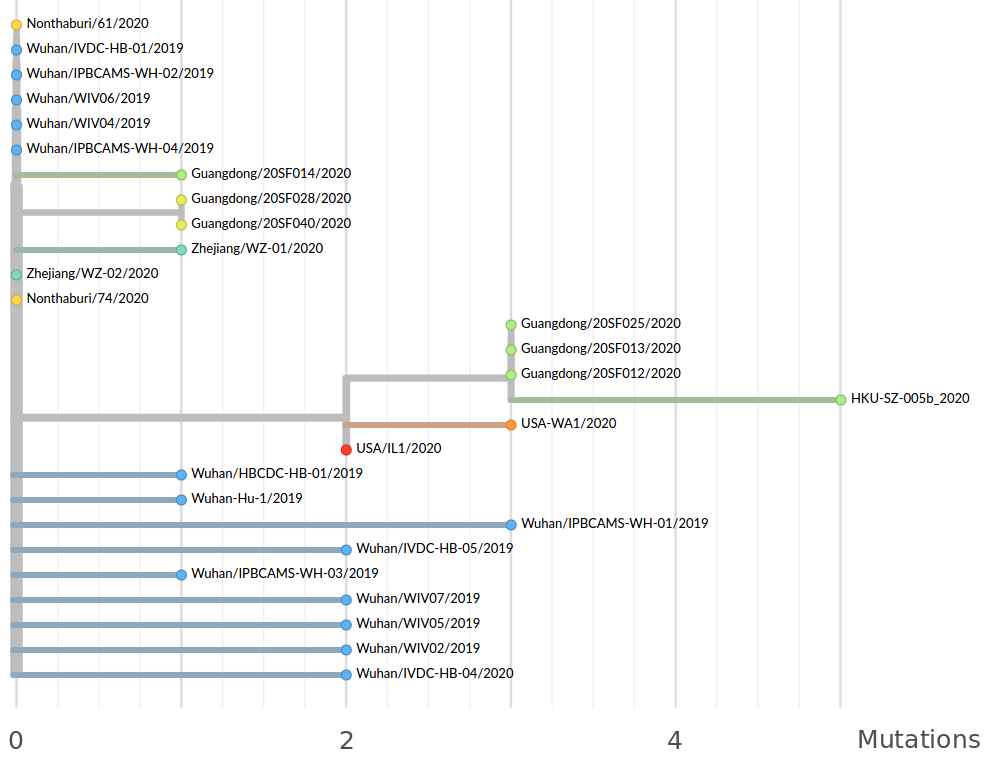
Early genomes differed by only a few mutations, suggesting very recent emergence
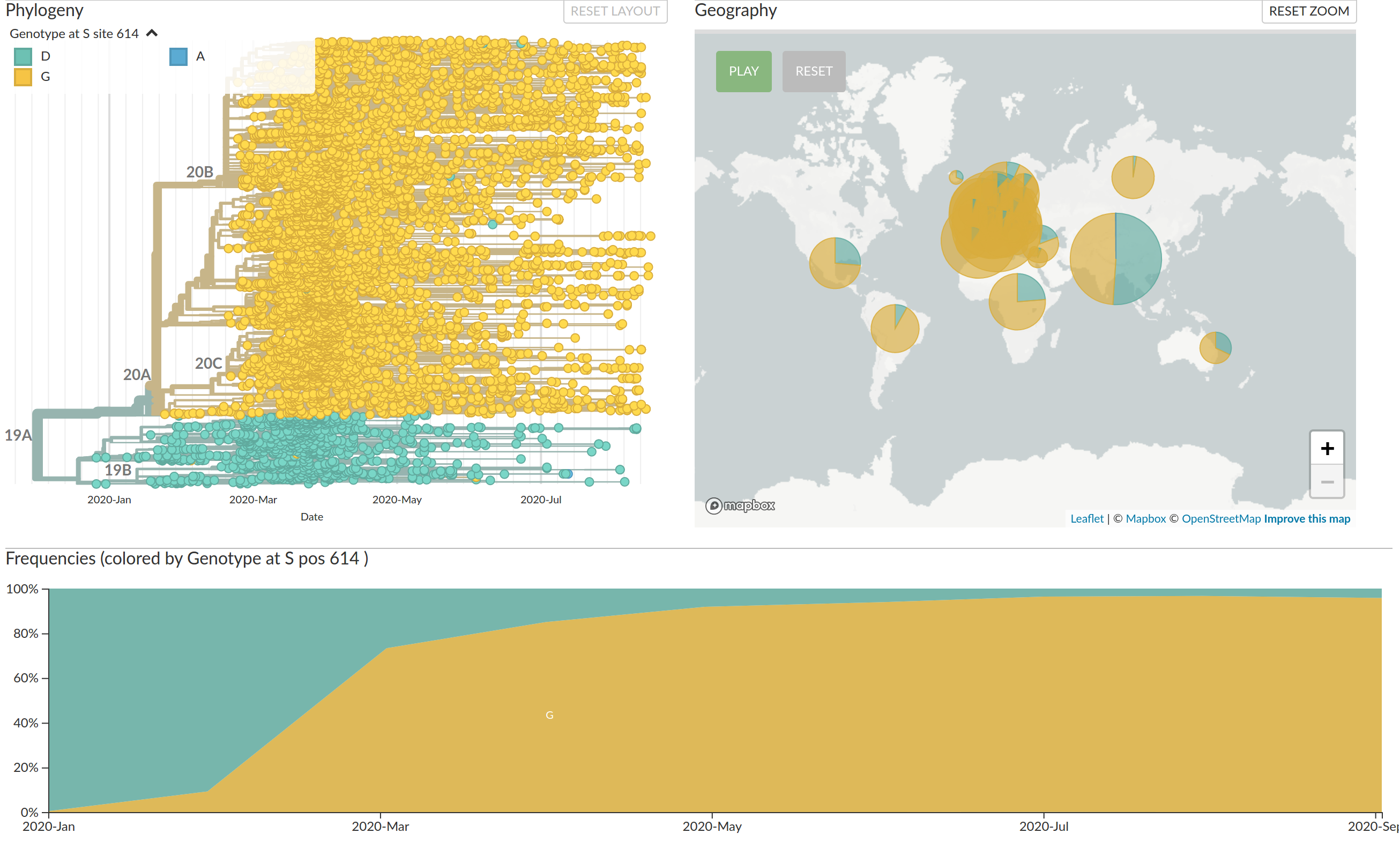
Emergence and dominance of VoCs
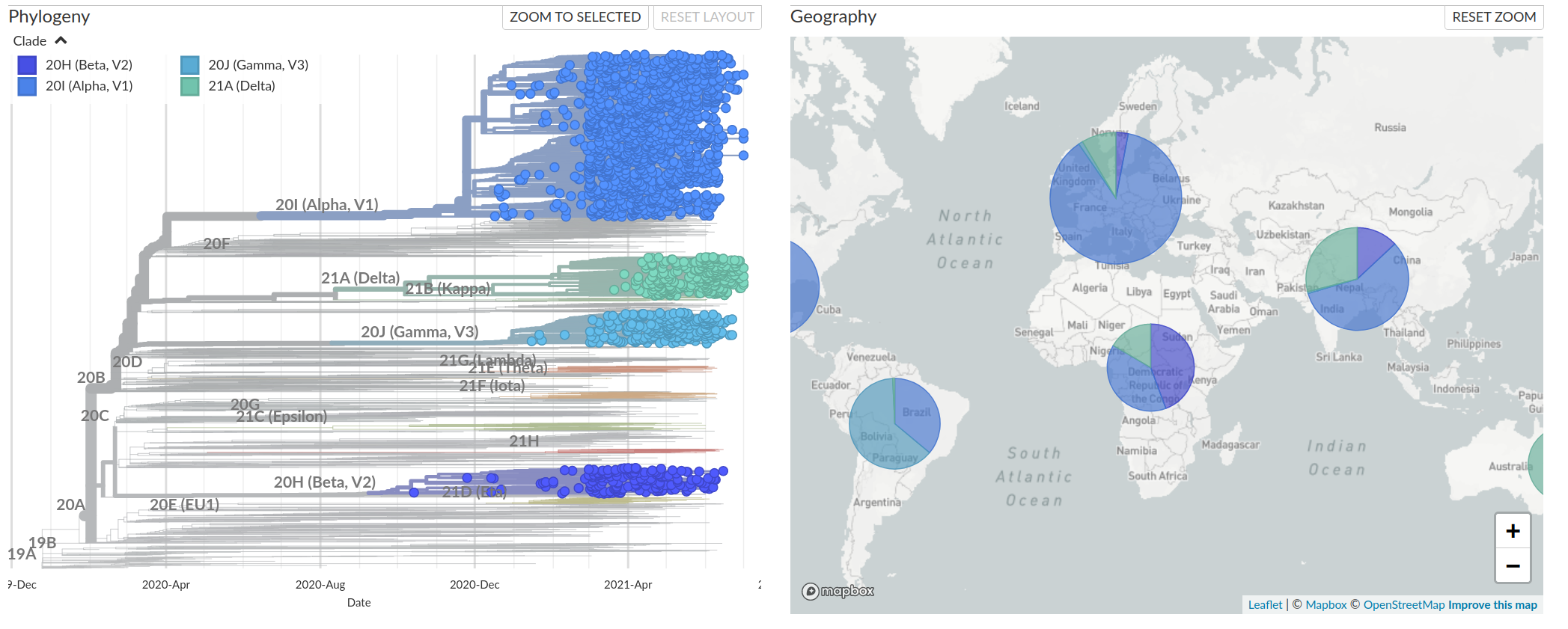
VoCs have more mutations than expected...
 nextstrain
nextstrain
VoCs and VoIs: rapid converging evolution
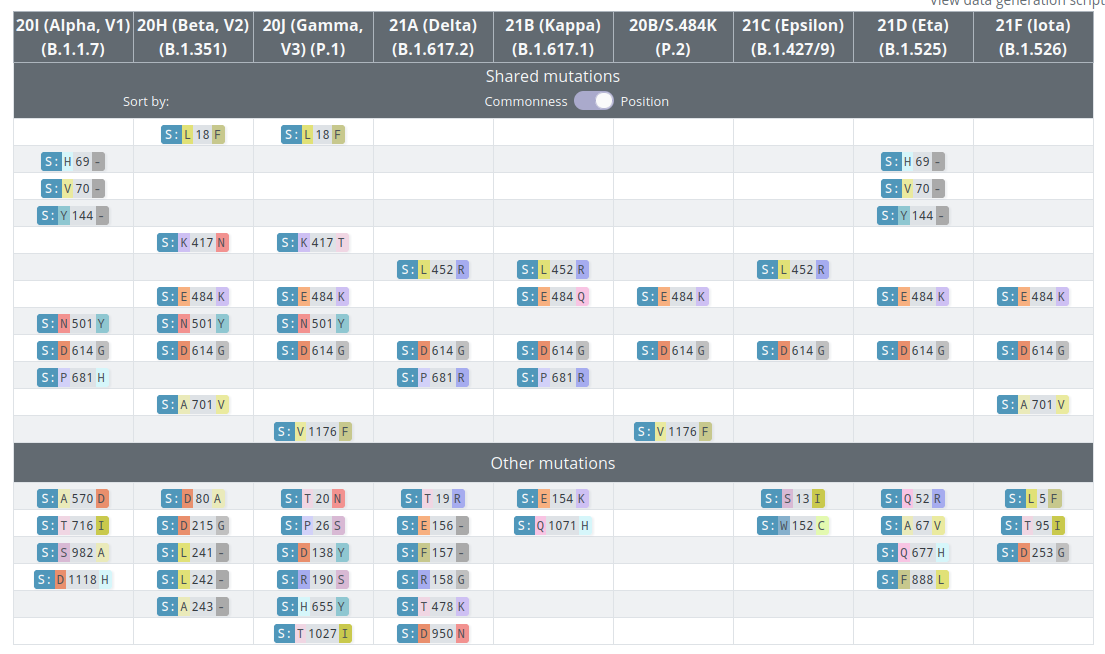 covariants.org by Emma Hodcroft
covariants.org by Emma Hodcroft
VoCs have been replacing other strains quickly
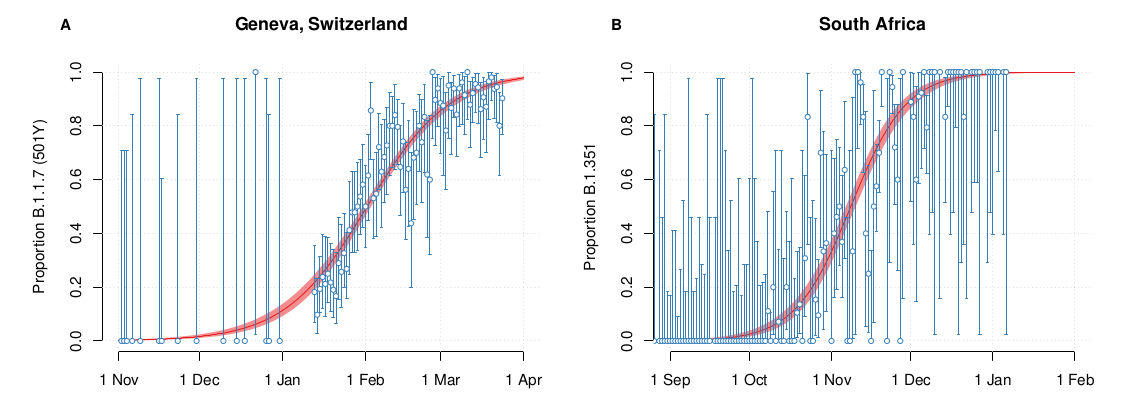
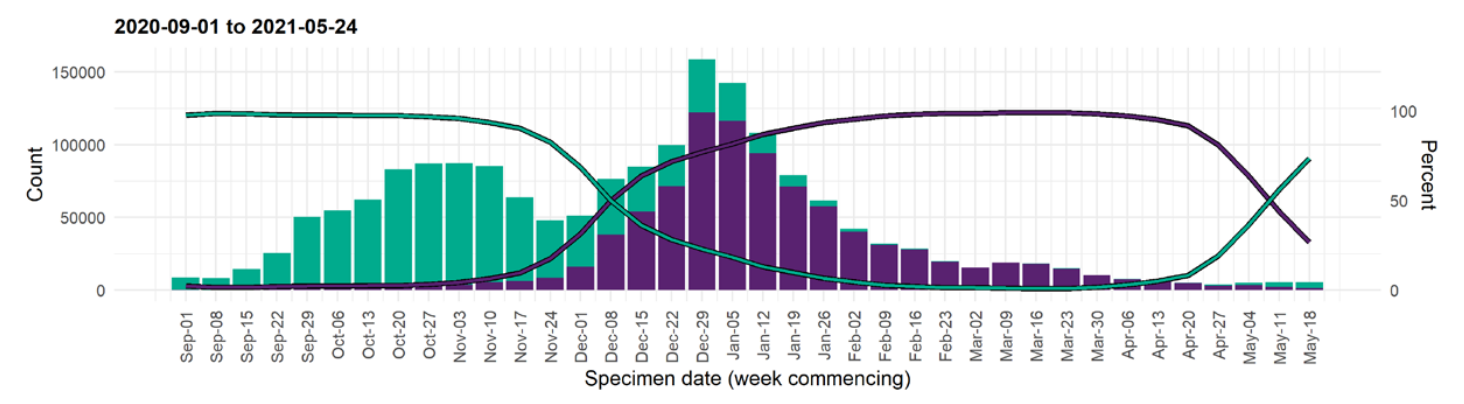 S-gene positivity in the UK
S-gene positivity in the UK
Relative growth advantage depends on setting
Africa
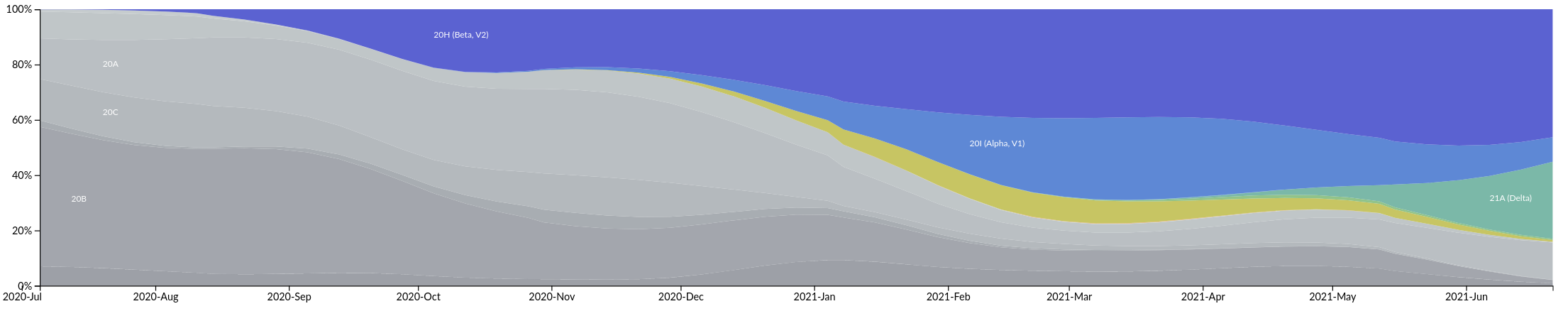
Asia
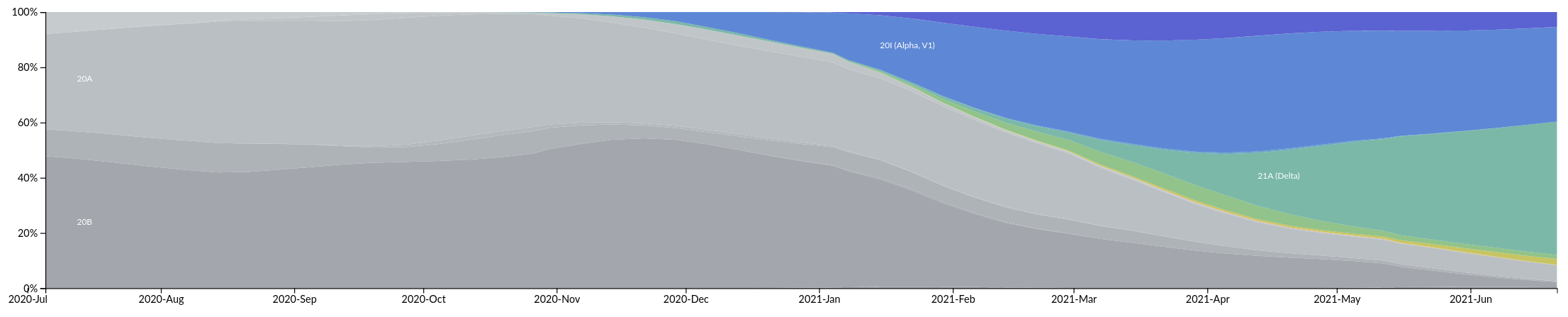
Europe
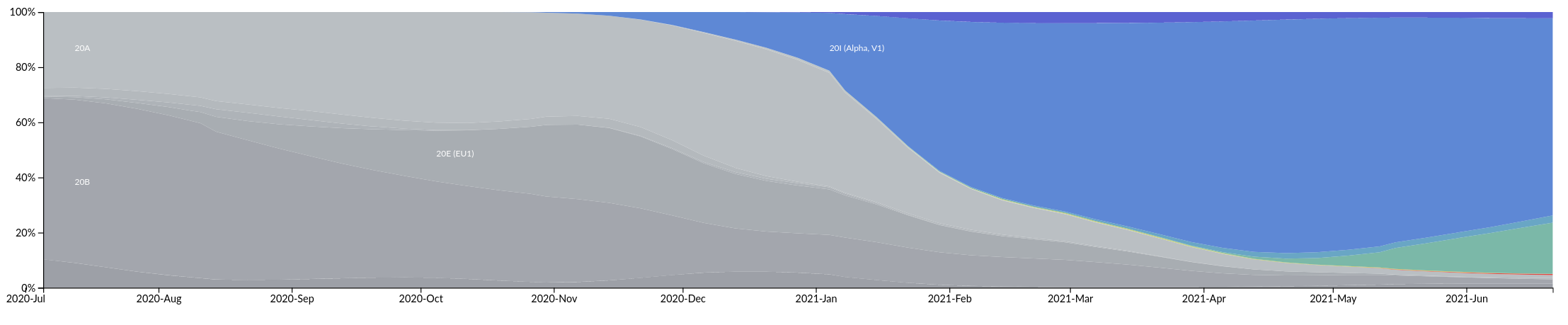

Asia

Europe

North America
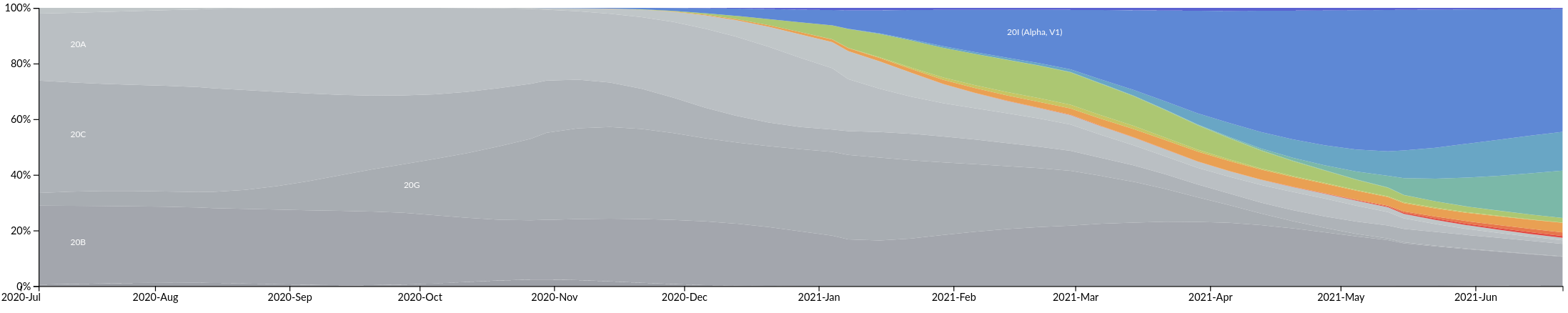
South America
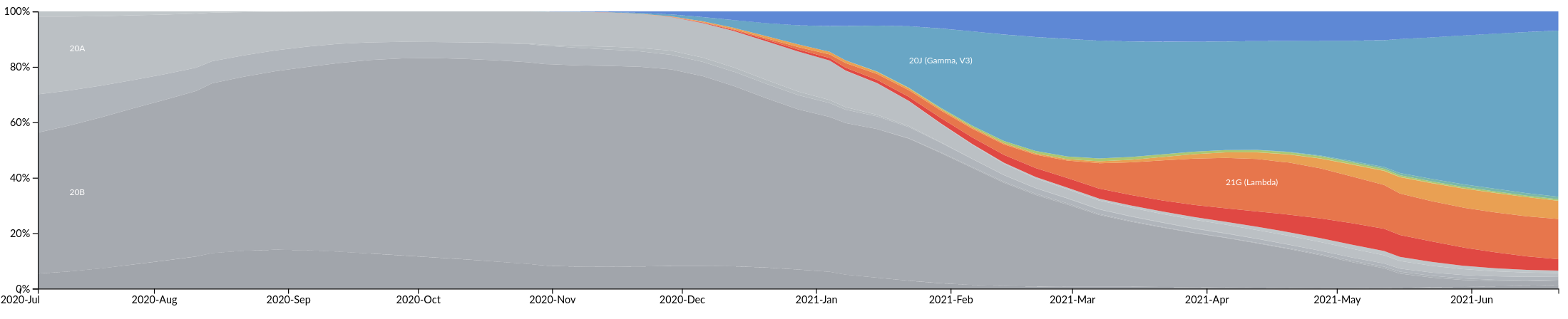

South America

- Intrinsic transmission advantage
- Immune evasion → more (mild) reinfections
- Likely depends on seropositivity in the population
- Dozens of other variants with comparably mutational signatures
How do we tell early on whether a variant is concerning?
How unusual are the patterns of convergent evolution?
SARS-CoV-2 variants can become dominant without transmission advantage
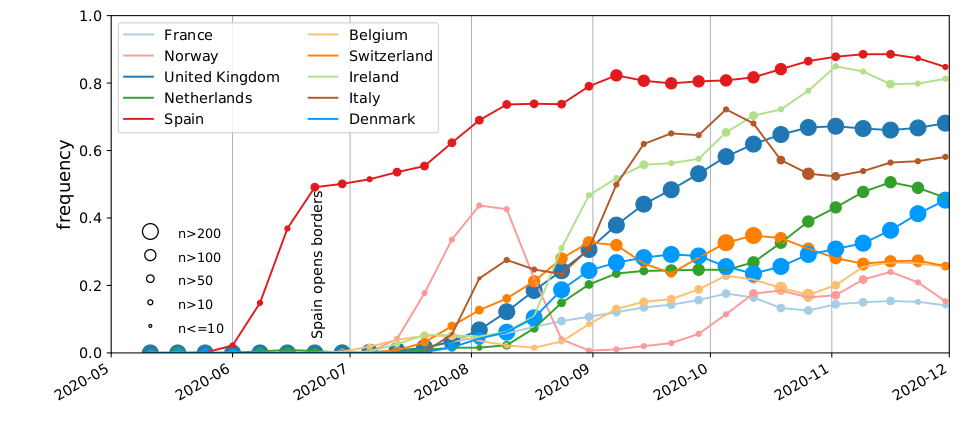
- Large differences in incidence (Spain vs most of Europe in Summer 2020)
- High travel volume during summer 2020
- Demographics with more social contacts
Convergent evolution is common in RNA viruses
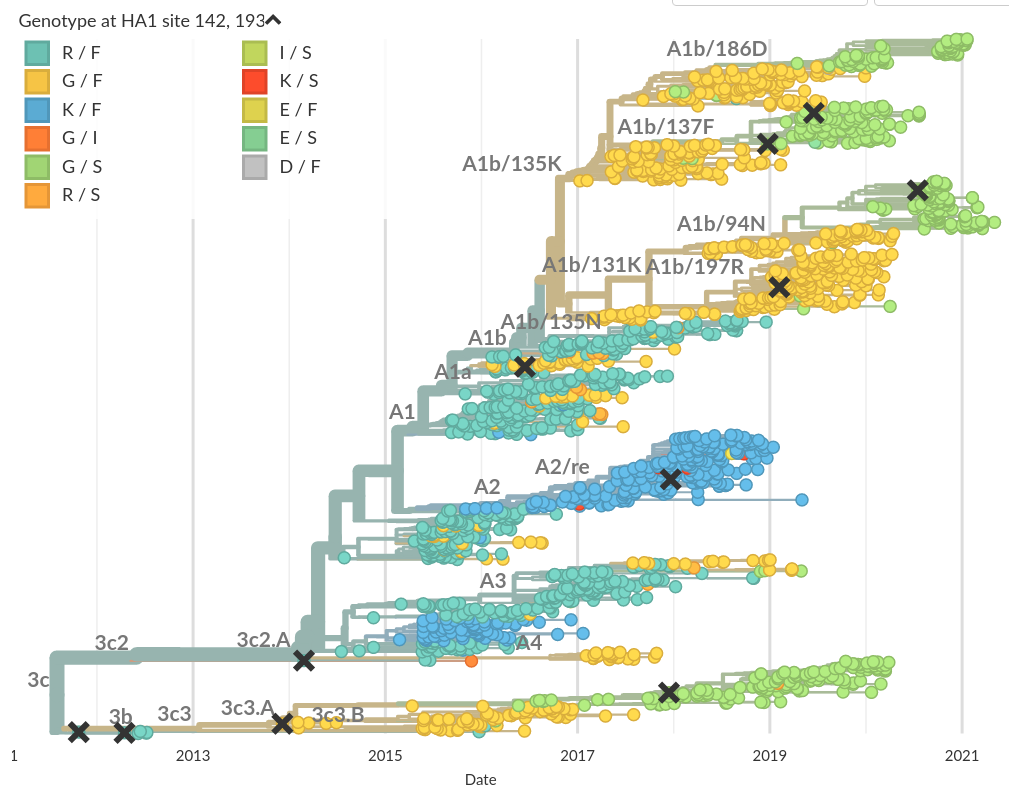
Recent Influenza A/H3N2 example:
4-fold convergence at positions 142 and 193 of HA1
- RNA viruses are not mutation limited
- In suitable environments, homoplasies are common
- Variation at homoplasic positions is often selected
→ but success depends on background
Mutational signatures
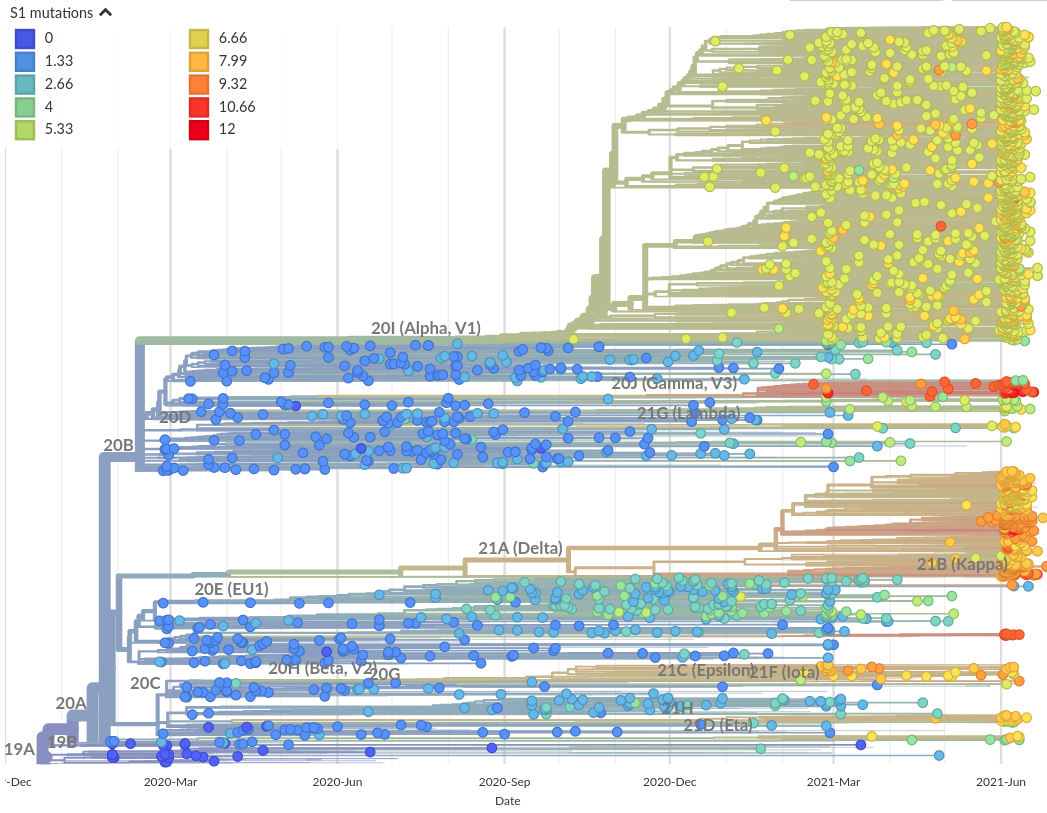
Number of amino acid changes in the S1 domain
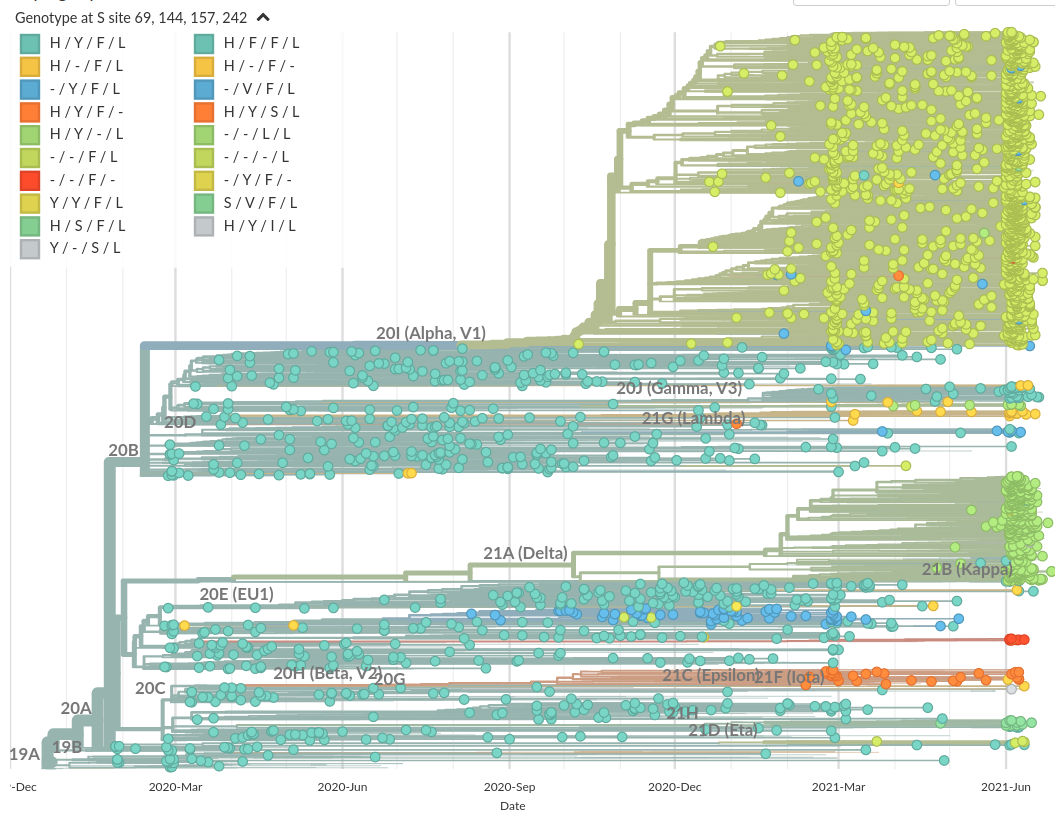
Prominent deletions the spike NTD
Common convergent deletion in NSP6
Growth measures and experimental scores
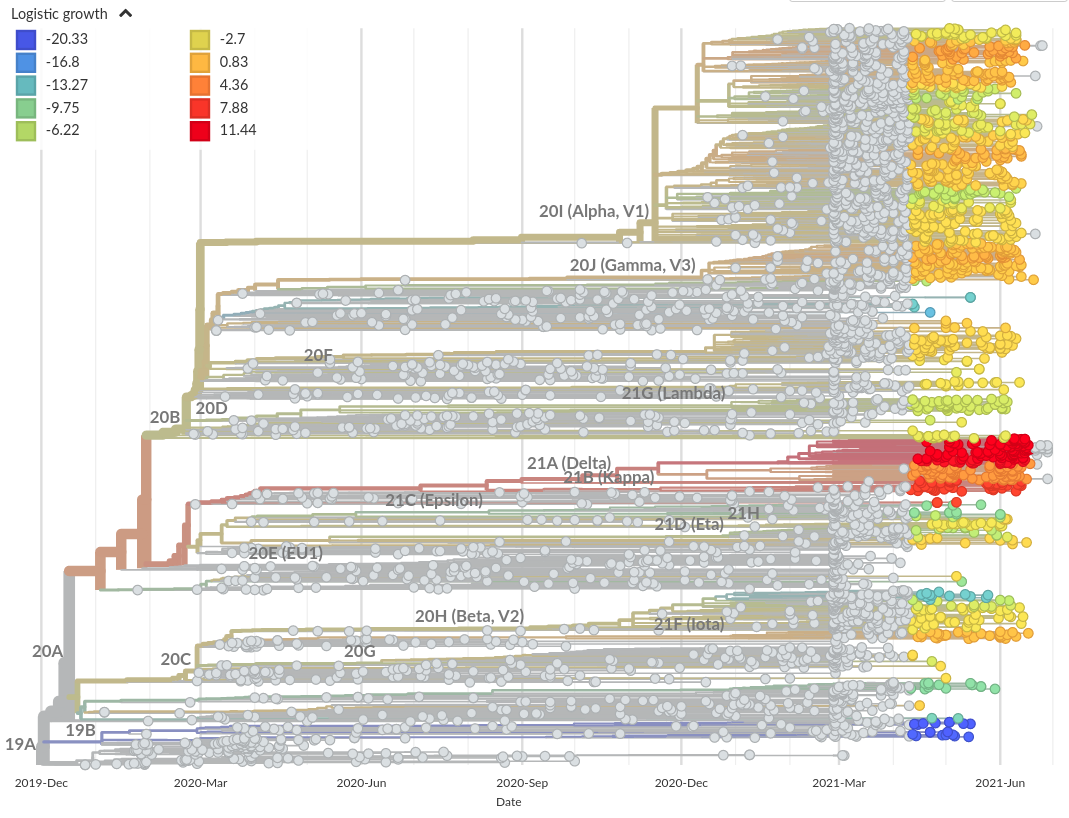
Recent growth rate
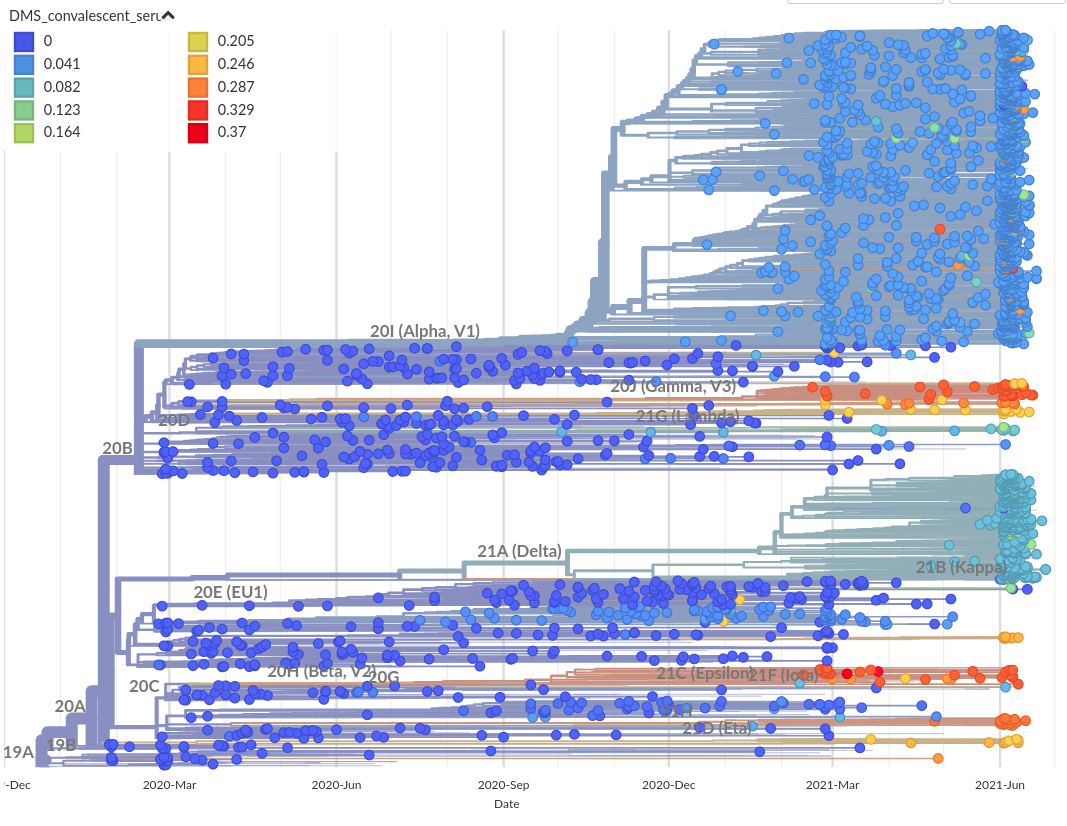
RBD escape based on deep mutational scanning.
Allie Greaney (Jesse Bloom's lab)
Conclusion
- Since late 2020, dominance of variants with abundant changes in Spike
- Success due to a complex mix of epidemiological, immunological and viral factors → hard to predict
- Regional variation in success -- alas VoC Delta is successful across the globe
- We don't know how flexible SARS-CoV-2 is...
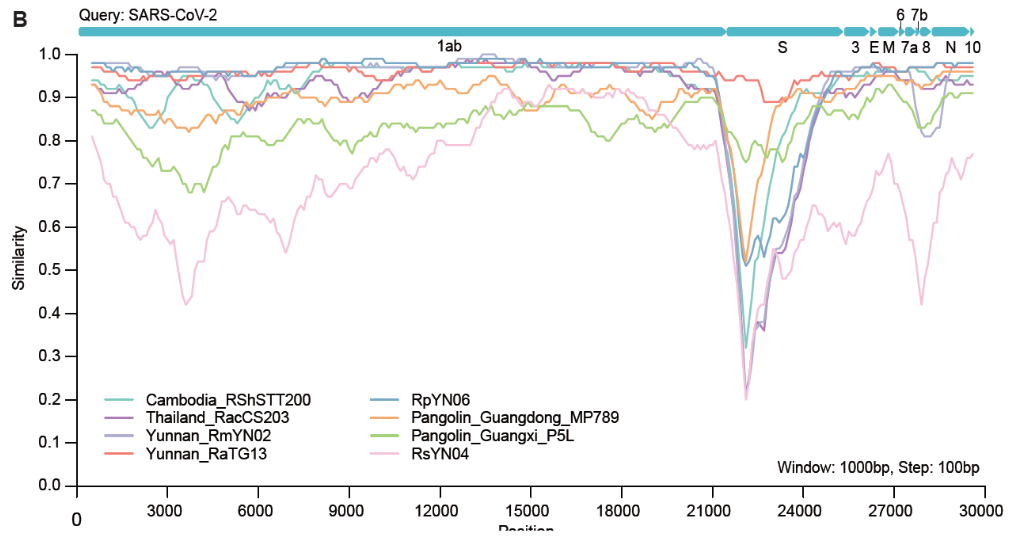 Zhou et al, 2021
Zhou et al, 2021
Acknowledgments
Trevor Bedford and his lab -- terrific collaboration since 2014

especially James Hadfield, Emma Hodcroft, Ivan Aksamentov, Moira Zuber, John Huddleston, and Tom Sibley
Data we analyze are contributed by scientists from all over the world
Data are shared and curated by GISAID

