Domestic animals are the primary source of animal-to-human transmission of influenza viruses and avian influenza is a major problem in poultry farming. For both of these reasons, influenza viruses in animals are closely monitored. Together with Trevor Bedford at the FHCRR in Seattle, our group has been developing automated phylogenetic analysis pipelines for Zika, Ebola, and human influenza viruses. These pipelines are rerun automatically as soon as new data are available and the results exposed via interactive web applications at nextstrain.org and nextflu.org.
In this project the current human influenza pipeline is to be expanded to animal influenza viruses. This will require adapting the phylogenetic analysis (written in python) to the more heterogenous and diverse animal data sets, and to adapt the visualization (written in js/d3) to the specifics of the animal influenza. In addition, phenotypic data such as antigenic properties have to be integrated with the genotypic data and interactions between different animal populations could be explicitly modeled.
As part of this project, the student will get familiar with and/or work on
- probabilistic phylogenetic inference
- ancestral state reconstruction
- integration of genotypic and phenotypic data
- data visualization d3/js
The project will be directly supervised by Richard Neher. Our interdisciplinary group combines methods and ideas from statistical physics with bioinformatics, population genetics, and phylogenetics. More information about our research, projects, and activities can be found at neherlab.org.
References and additional information
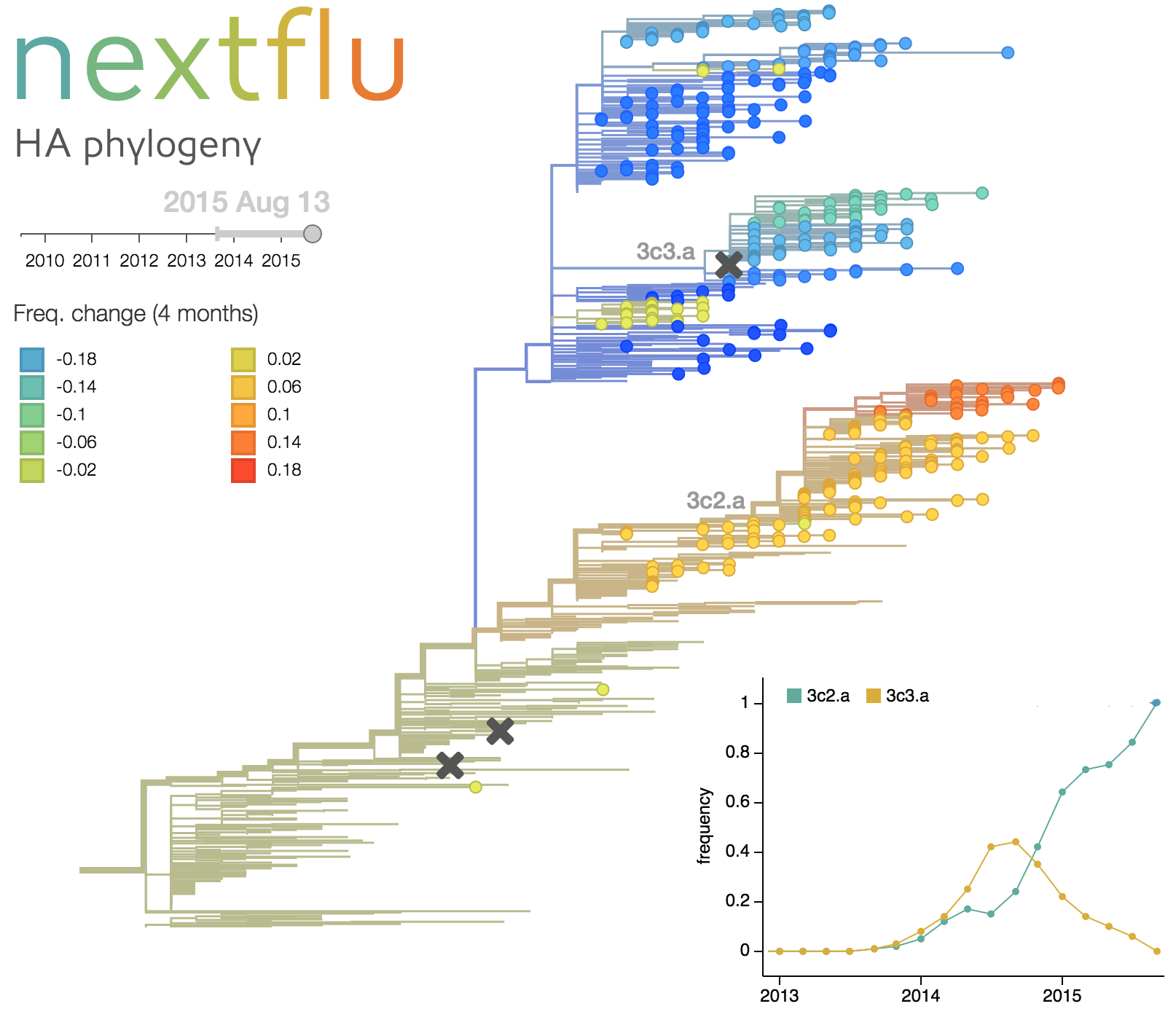
- Real-time tracking of seasonal influenza at nextflu.org
- Lab website: neherlab.org
- Existing code base: github.com/nextstrain
- Richard A. Neher and Trevor Bedford. Bioinformatics (Oxford, England), vol. 31, 3546--3548, 2015